This article shows the excellent sensitivity and precision of the SPEAR UltraDetect platform, demonstrated by two SPEAR UltraDetect™ assays targeting critical neurological biomarkers: Glial Fibrillary Acidic Protein (GFAP) and Neurofilament-Light (NF-L).
These assays provide ultra-sensitivity with high precision, allowing for the reliable identification of subtle, clinically relevant changes that other ultra-sensitive platforms frequently miss due to their high variability.
With impressively low coefficients of variation (CVs) even at low concentrations, SPEAR establishes a new standard for analytical performance, making it an effective instrument for early identification and longitudinal investigation of neurological disorders.
Introduction
Accurate biomarker measurement, combining high sensitivity and precision, is critical for neurological research and therapeutic applications. Biomarkers like as GFAP, which indicates astrocytic activation, and NF-L, which reflects axonal damage, are present at low levels in the blood early in the disease and can rise gradually over time.1-3
Quantifying these indicators at very low levels in illnesses like Alzheimer's disease, multiple sclerosis, and stroke allows for better disease management and faster intervention.
This necessitates the detection of low-abundance biomarkers and their consistency and reproducibility across time and samples. The ability to identify disease-related alterations from assay variability requires high precision in biomarker measurements.
While conventional ultra-sensitive systems have enabled novel discoveries, they have always struggled with assay precision and variability, particularly in blood matrices, where neurobiomarker concentrations are often much lower than in CSF.
These constraints compromise the proper tracking of small but significant biomarker variations, which are critical for recognizing early disease signals, assessing therapy response, and tracking longitudinal disease development.
Successive Proximity Extension Amplification Reaction (SPEAR) is a homogenous, ultrasensitive immunoassay method that addresses the sensitivity limitations and complexity of traditional immunoassays.
SPEAR uses a novel two-factor authentication approach to ensure that amplifiable signals are only created through prolonged co-localization of probes on target proteins.
It accurately measures protein biomarkers at attomolar concentrations with as little as 1 µl of diluted sample, outperforming the most sensitive heterogeneous platforms on the market.
SPEAR's homogeneous nature eliminates nonspecific binding associated with solid-phase capture, giving it higher specificity than heterogeneous immunoassay platforms. SPEAR's procedure eliminates error-prone wash phases, resulting in extremely high precision.
SPEAR UltraDetect™ assays, based on this core technology, provide unmatched sensitivity for measuring low-abundance biomarkers while maintaining high specificity and precision.
SPEAR UltraDetect reads data using standard qPCR tools, resulting in extremely consistent results across multiple qPCR platforms and formats. It is simple to integrate into existing laboratory settings and offers exceptional scalability.
This article provides results demonstrating the outstanding precision of the SPEAR UltraDetect GFAP and NF-L assays, 100 % detectability and quantifiability in healthy samples, and excellent agreement with the GFAP and NF-L tests on the widely used Simoa platform.
Materials and methods
Samples from healthy donors and diseased individuals were obtained from multiple commercial sources, such as Precision for Medicine and BIOIVT, and tested using SPEAR UltraDetect™ NF-L (item 90004, Spear Bio) and GFAP (item 90010, Spear Bio), as well as Simoa® NF-Light® Advantage PLUS (item 104364, Quanterix) and Simoa GFAP Advantage PLUS (item 104619, Quanterix).
SPEAR UltraDetect™ method
Samples were handled using the Formulatrix® F.A.S.T™ liquid handler and QInstruments® BioShake® iQ, as instructed by SpearBio. To begin, combine 1 µL of probe mix with 1 µL of sample or calibrator, then incubate for 1 hour.
After incubating 6 µL of reaction mix for 20 minutes, 12 µL of qPCR mix was added for final analysis. The qPCR findings were translated to protein concentrations using the SPEARview analysis software. (Figure 1).
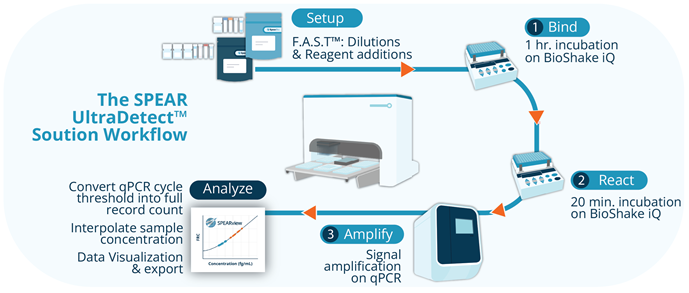
Figure 1. The full SPEAR UltraDetect Assay workflow. Image Credit: Spear Bio
Quanterix Simoa method
Samples were run on the Simoa HD-X Analyzer® according to the manufacturer's instructions.
To begin, 100 µL of sample or calibrator was mixed with 25 µL bead reagent, followed by washing (3x), addition of 100 µL detection reagent, washing (3x), addition of 100 µL SBG reagent, washing (4x), and addition of 50 µL RGP. Protein measurements were generated using the Quanterix software.
Results
Precision
Intra-plate CVs for both GFAP and NF-L were consistently less than 10 %, with average CVs of 3.0 % and 3.2 %, respectively (Figure 1A, 2A), demonstrating greater precision compared to previously published Simoa assays.4,5
Sensitivity
SPEAR UltraDetect detects GFAP and Nf-L at pg/mL (Table 1), identifying and quantifying 100 % of healthy samples, with the lowest endogenous readings across all investigated matrices at least one order of magnitude above the functional LLoQ (Figure 1B, 2B).
This level of sensitivity implies that baseline and minor but disease-relevant changes in GFAP and NF-L can be accurately quantified in a wide range of neurology research applications.
Concordance
Measurements of GFAP and NF-L from SPEAR UltraDetect and Simoa are extremely consistent across healthy and sick samples, with R2 values of 0.941 and 0.996, respectively. (Figure 3).
Table 1. SPEAR UltraDetectTM NF-L and GFAP Assay specifications in EDTA plasma and serum samples. Source: Spear Bio
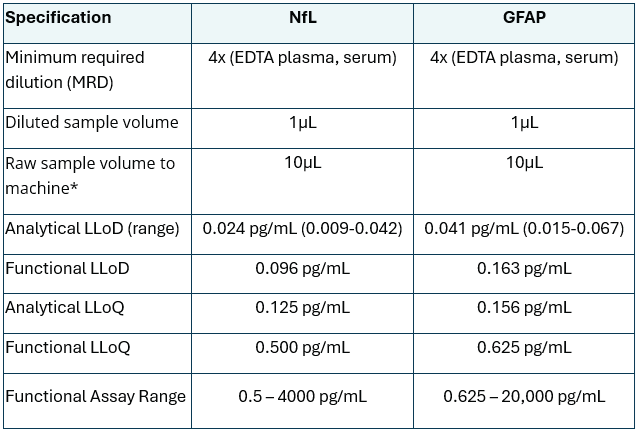
* Can be used for multiple replicates and biomarker measurements.
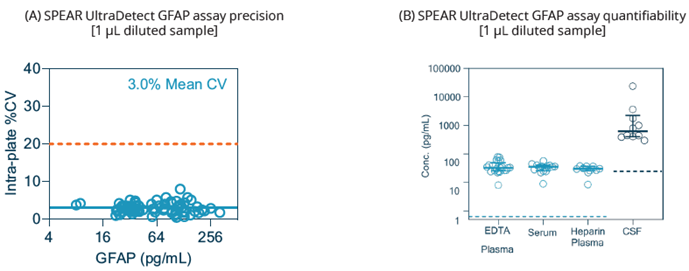
Figure 2. Precision and quantifiability of SPEAR UltraDetect GFAP assay. (A) Intra-plate CV plot from a total of 70 samples, including matched EDTA plasma (n=15) and serum
(n=15) healthy controls, amyloid negative plasma (n=6), and amyloid positive plasma (n=34). The 20 % CV reference line is marked; (B) Endogenous readings of human EDTA plasma. *Heparin plasma and CSF samples are tested for reference only. Image Credit: Spear Bio
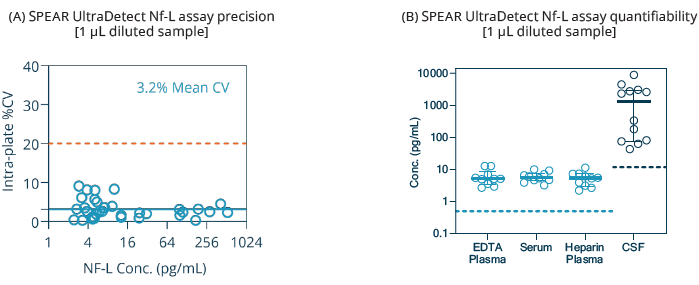
Figure 3. Precision and quantifiability of SPEAR UltraDetect NF-L assay. (A) Intra-plate CV plot from a total of 34 samples, including matched EDTA plasma (n=10) and serum
(n=10), and amyotrophic lateral sclerosis (n=14). The 20 % CV reference line is marked; (B) Endogenous reading of human EDTA plasma (n=10), serum (n=10), heparin plasma* (n=10), and CSF* (n=10). Functional LLoQ is indicated as dashed lines. Bars indicate the mean with the interquartile range. *Heparin plasma and CSF samples are tested for reference only. Image Credit: Spear Bio
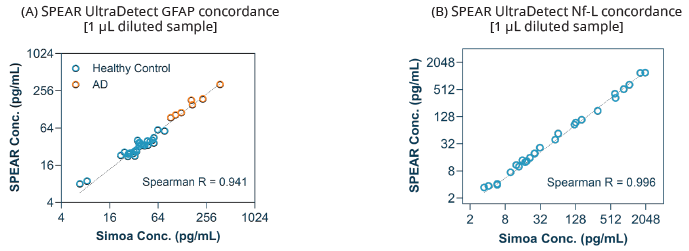
Figure 4. Assay concordance with Simoa platform. (A) 37 samples, including matched EDTA plasma (n=15) and serum (n=15) samples from healthy controls, and 7 AD
donors were measured using SPEAR UltraDetect GFPA and Simoa GFAP Advantage PLUS; (B) 26 plasma samples, including healthy controls (n=8), multiple sclerosis (MS, n=6), AD (n=2), progressive supranuclear palsy (PSP, n=3), amyotrophic lateral sclerosis (ALS, n=1), PSP/ALS spiked (n=6), were measured using SPEAR UltraDetect NF-L and Simoa NF-Light Advantage PLUS. Image Credit: Spear Bio
Discussion
The remarkable sensitivity and precision of SPEAR UltraDetect, as demonstrated in this article, are significant advantages for detecting early, subtle changes in neurodegenerative biomarkers.
SPEAR UltraDetect enables researchers to identify GFAP and NF-L at ultralow levels with high precision, allowing them to record the earliest shift in protein biomarker levels in prodromal and mild stages of neurological disorders.
In clinical research, SPEAR UltraDetect's high precision makes it easier to detect and follow biomarker changes as disease progresses or in response to treatment, allowing for powerful biomarker monitoring in longitudinal studies.
SPEAR's demonstrated combination of high sensitivity, precision, and scalable implementation establishes it as a game changer for neurology applications ranging from early disease identification to treatment development.
References
- Wang, X., et al. (2024). Peripheral GFAP and NfL as early biomarkers for dementia: longitudinal insights from the UK Biobank. BMC medicine, 22(1). DOI: 10.1186/s12916-024-03418-8. https://link.springer.com/article/10.1186/s12916-024-03418-8.
- Samadzadeh, S. and Sleator, R.D. (2025). The role of Neurofilament light (NfL) and glial fibrillary acidic protein (GFAP) in MS and AQP4-NMOSD: Advancing clinical applications. eNeurologicalSci, [online] 38, p.100550. DOI: 10.1016/j.ensci.2025.100550. https://www.sciencedirect.com/science/article/pii/S2405650225000048?via%3Dihub.
- Barro, C., et al. (2022). Serum GFAP and NfL Levels Differentiate Subsequent Progression and Disease Activity in Patients With Progressive Multiple Sclerosis. Neuroimmunology and Neuroinflammation, [online] 10(1), pp.e200052–e200052. DOI: 10.1212/nxi.0000000000200052. https://www.neurology.org/doi/10.1212/NXI.0000000000200052.
- Simoa GFAP Advantage PLUS data sheet (available at www.quanterix.com)
- Simoa NF-L Advantage PLUS data sheet (available at www.quanterix.com)
About Spear Bio
Spear Bio is an innovative leader in providing scalable solutions for ultra-sensitive protein biomarker measurements. Spear Bio’s proprietary technology, Successive Proximity Extension Amplification Reaction (SPEAR), employs a unique 2-factor authentication mechanism to precisely measure protein biomarkers at attomolar level from sub-microliter sample volume. Spear Bio is focused on leveraging its technology’s unprecedented sensitivity to transform protein research and early disease diagnosis.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.net, which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.