Since the beginning of the coronavirus disease 2019 (COVID 19) pandemic, it has been established that the causative virus, severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) gains entry to the target host cell via its spike glycoprotein.
Many mutations have since occurred in this viral protein, some with the ability to alter the antibody recognition site. A recent paper, published as a preprint on the bioRxiv* server, reports on a pair of linked mutations, ΔH69/ΔV70, which has shown repeated emergence and transmission.
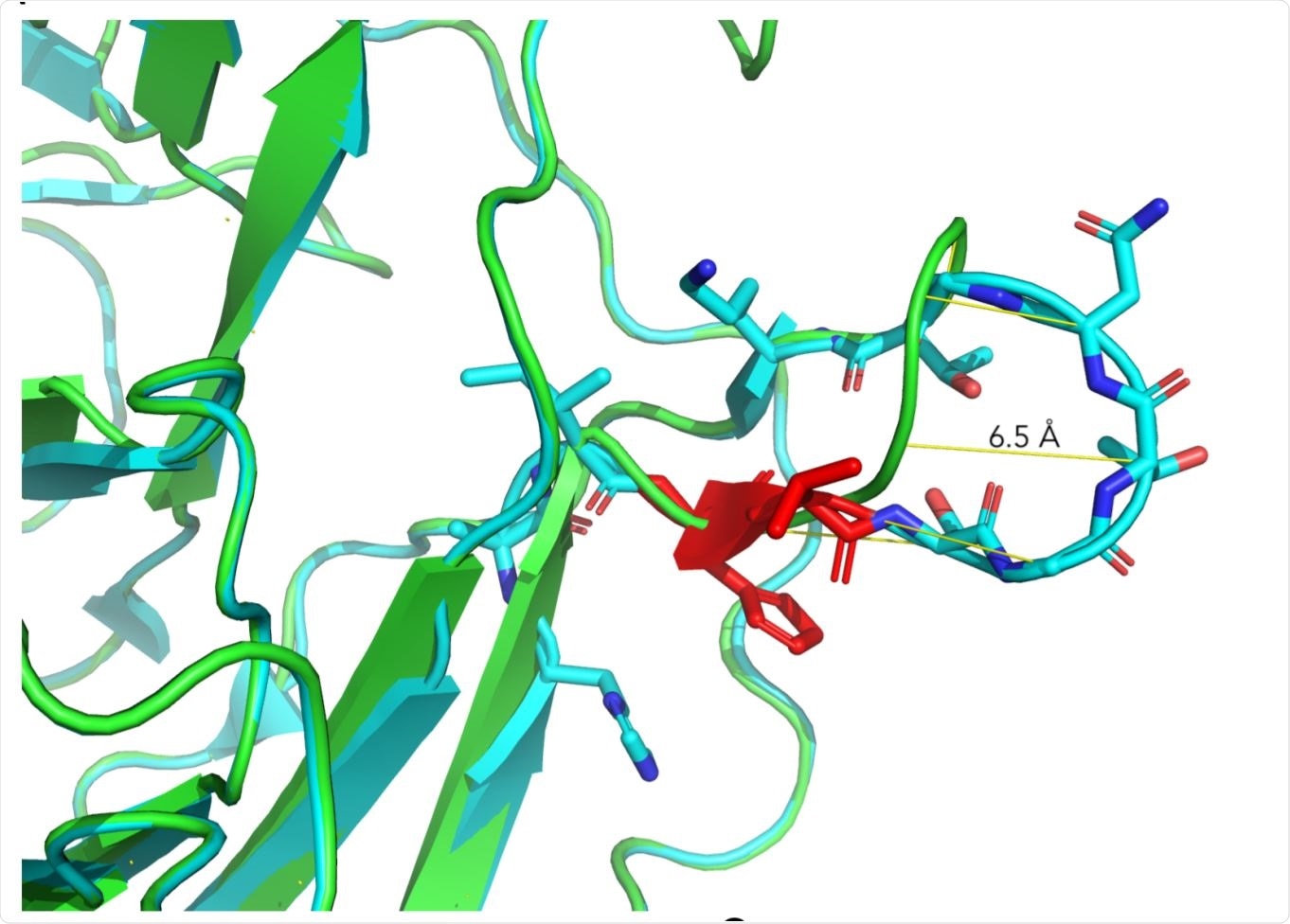
Structural aspects of ΔH69/V70. Image Credit: https://www.biorxiv.org/content/10.1101/2020.12.14.422555v2.full.pdf

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Escape mutations emerge following plasma use
The viral spike protein attaches to the host cell receptor, angiotensin-converting enzyme 2 (ACE2), found in many human tissues. Though the spike protein is central to viral entry, its receptor-binding domain (RBD) has remarkable mutational tolerance.
The increasing use of CP has led to the emergence of spike ΔH69/ΔV70 and D796H variants. These mutations in the spike N-terminal domain (NTD) may allow escape from neutralizing anti-NTD antibodies in CP but without impact on infectivity.
Tracking the spike ΔH69/ΔV70 mutation
With the enormous scale on which SARS-CoV-2 genome sequencing is occurring at present, it has become possible to examine multiple parameters for their correlation with increased infectivity and transmissibility.
The current study aimed to explore the underlying situation in which the ΔH69/ΔV70 mutation arose, using data from the Global Initiative on Sharing All Influenza Data (GISAID) data.
They found that of the available data, about 2.5%, or 3,000 sequences, contained this deletion, though most of them were necessarily from Europe since this was the origin of the majority of GISAID sequences. The presence of the mutation in many different lineages reflected its independent emergence at least five times.
First occurring independently in Thailand and Germany, as far back as January 2020, its prevalence went up after a few months, from August 2020. In some variants, this double mutation was present in isolation, and in others, they were present along with other RBD mutations.
Modeling mutation effects
The researchers created a homology model of the spike NTD with the double mutation. They observed that in this model, the mutation led to a shift in the positions of several residues, resulting in a predicted change in the conformation of a protruding loop made up of amino acids from positions 69-76, by pulling it into the NTD.
Independent emergence of mutation sets
They looked at viral lineages with a high frequency of spike mutations, especially in combination with N439K. The latter may prevent neutralization by some anti-RBD monoclonal antibodies (mAbs), despite showing an intact affinity for ACE2. The frequency of the double mutation increased markedly from August onwards, in combination with N439K. By November, the number of sequences with all three mutations was double that of sequences containing only N439K.
Another cluster showed the double deletion emerging in a lineage containing Y453F, an RBD mutation that resulted in increased binding affinity for the ACE2 receptor. This was again associated with reduced neutralization by COVID-19 antiserum. This has been found only in Danish sequences so far.
Thirdly, another RBD mutation N501Y has been associated with the same double deletion. Its location raises the possibility that it may be an escape mutation from antibodies like CoV2-2499. Moreover, the passage of SARS-CoV-2 in mice led to the emergence of the single mutation with an increased pathogenic effect.
Sequences with N501Y alone have been found in the UK and Brazil from April onwards. However, the number of sequences with N501Y along with the double deletion increased following their emergence in the UK in September 2020. At present, they exceed the number of sequences containing the single mutation.
This especially relates to a sub-lineage containing ~350 sequences, with six novel mutations involving the RBD (including N501Y), spike S2 subunit, and the double deletion. This long-branch mutation set could either have come from an area with very little surveillance or from a chronically infected individual, where the virus persisted and replicated in the host for a long period.
What are the implications?
Multiple clusters of virus variants have emerged in several episodes. In all these cases, the double deletion has been found to emerge in association with a mutation of the RBD that reduces receptor binding by mAbs and is increasing in prevalence. NTD deletions have the potential to increase viral transmissibility.
The double deletion affecting an exposed surface which may enable the variant to escape immune recognition. This could be by shifting linkages between the spike subunits and altering RBD orientation, thus impacting viral spread and infectivity.
The only measures to prevent such escape mutations from hampering containment strategies are to take increased care of non-pharmaceutical interventions and speed up vaccination coverage to herd immunity levels.
The development of rapid diagnostic tests for the double deletion will be important in signaling the emergence of antibody escape mutations and thus enabling global monitoring of virus adaptation.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Kemp, S. A. et al. (2020). Recurrent emergence and transmission of a SARS-CoV-2 Spike deletion ΔH69/ΔV70. bioRxiv preprint. doi: https://doi.org/10.1101/2020.12.14.422555. https://www.biorxiv.org/content/10.1101/2020.12.14.422555v2
- Peer reviewed and published scientific report.
Meng, Bo, Steven A. Kemp, Guido Papa, Rawlings Datir, Isabella A.T.M. Ferreira, Sara Marelli, William T. Harvey, et al. 2021. “Recurrent Emergence of SARS-CoV-2 Spike Deletion H69/V70 and Its Role in the Alpha Variant B.1.1.7.” Cell Reports, June, 109292. https://doi.org/10.1016/j.celrep.2021.109292. https://www.cell.com/cell-reports/fulltext/S2211-1247(21)00663-X.