Scientists have developed several coronavirus disease 2019 (COVID-19) vaccines, which received approval or emergency use authorization (EUA) from global regulatory bodies to manage severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) infection.
SARS-CoV-2 is the causal agent of the ongoing COVID-19 pandemic that has claimed more than 4.8 million lives worldwide. Interestingly, previous studies have indicated that both humoral (antibody-mediated) and cell-mediated immunity is essential to completely clear SARS-CoV-2 infection.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
B cell Depleting Therapies and SARS-CoV-2 Infection
Immunocompromised patients subjected to B-cell depleting therapies were found to experience prolonged SARS-CoV-2 infection and viral shedding. Although viral shedding could occur for months after the onset of symptoms, viable viruses have not been detected after nine days post-illness symptoms. Persistent viral replication may cause profound lymphocyte defects owing to B- and T-cell depleting therapies or underlying hematologic disease. An important aspect of viral persistence in immunosuppressed settings is the development of variants, especially during convalescent plasma therapy.
Several recently published studies have revealed that SARS-CoV-2 in immunocompromised hosts tends to cause deletion mutations in the spike protein, especially in the S1 domain. Generally, the S1 region of the spike protein of SARS-CoV-2 is responsible for establishing the host-virus interaction to promote viral invasion to the host cell. In both immunocompetent and immunosuppressed infected hosts, deletions may occur across the viral genome that can alter the mechanism of the virus-host interaction. Researchers found that accessory proteins such as ORF7ab and ORF8, which are associated with early stimulation of IFN-γ production, were deleted in a large number of immunocompetent individuals.
A New Study
A new study published on the medRxiv* preprint server focused on a patient with a persistent symptomatic viral infection for over a year. Researchers amplified sub-genomic transcripts for the E gene (sgE) using nasopharyngeal samples of the patient infected with SARS-CoV-2 over the course of 335 days. This patient also had a history of CAR T-cell therapy. The SARS-CoV-2 tests had been carried out using the nasopharyngeal swabs of the patient, and, interestingly, they remained COVID-19 positive persistently. Complete genomic sequencing was conducted on nasopharyngeal samples from the patient.
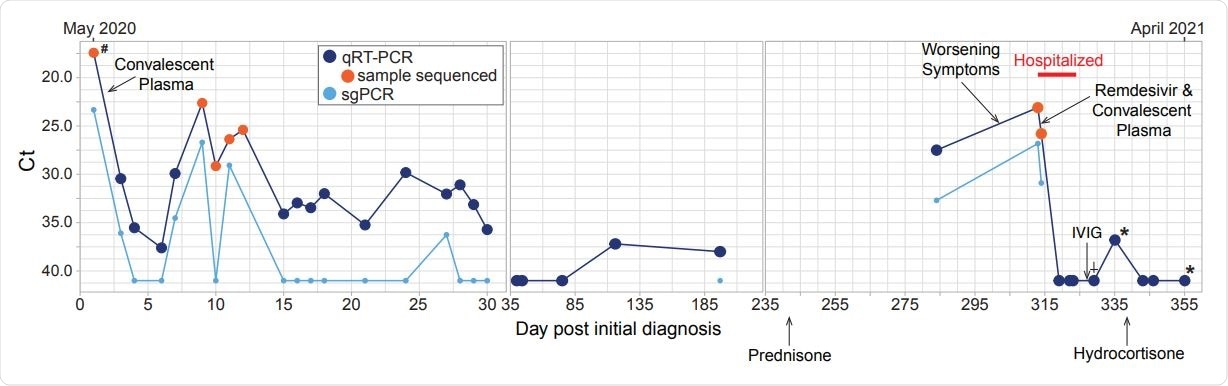
Timeline of diagnostic tests for SARS-CoV-2 and treatment. Nasopharyngeal or oropharyngeal (upper respiratory) specimens were collected for detection of SARS-CoV-2 RNA, except when indicated by the following: # indicates BAL sample, * indicates days when sputum specimens were collected and † indicates days when saliva was collected. Specimens with Ct values over 40 were considered negative for SARS-CoV-2 RNA. PCR for sub-genomic RNA was performed only on specimens that tested positive for genomic RNA. Samples that were used for next-generation sequencing are indicated with an orange circle. Treatments administered are indicated with a black arrow and labeled.
The Main Findings
Analysis of the sequencing data revealed persistence of COVID-19 infection over a 335-day period. The authors stated that to date, this is perhaps the most prolonged reported infection with SARS-CoV-2. This study indicated an evolutionary trajectory of the virus based on the mutation rate of the polymerase. Researchers found that the patient exhibited SARS-CoV-2 symptoms for the entire period of infection.
The current studies showed that the virus accumulated a unique in-frame deletion in the amino-terminal domain (NTD) of the spike protein, over the period of a year. A similar kind of deletion had been observed in the Lambda variant (B.1.351), which contained NTD deletion and an R246I mutation. The reason why deletion in that region of the NTD occurs in an immunocompromised patient is not yet apparent. However, this report is in line with previous studies showing chronic SARS-CoV-2 infection occurs in severely immunocompromised hosts who underwent convalescent plasma treatment. Also, this condition leads to the emergence of SARS-CoV-2 variants and reduced effectiveness of neutralizing antibodies. The immunocompromised patient considered in this study was also subjected to a convalescent plasma treatment during her first hospital admission.
This study also reported 497nt deletion in the ORF7b and ORF8 genes, and it is the longest deletion reported in this region of the genome. Such deletion has been observed for the first time in an immunocompromised patient. A previous study had reported ORF7b/ORF8 deletions in this region span from 62nt to 382nt. This mutation was first identified in Singapore in January of 2020.
Prior studies have indicated that deletion mutants may be less effective in establishing infection in a new host owing to the loss of immune evasion features of ORF8. Earlier studies have also shown that ORF8 is a key antagonist of innate immunity. Therefore deletion of this gene elicits a robust and highly specific immune response during infection. Interestingly, a large deletion was found in the immunocompromised patient, which might have occurred because the immunocompromised nature of this patient eliminated a need for ORF8 during infection.
The authors reported that when the patient was treated with remdesivir and high-titer convalescent plasma, after the appearance of deletion mutations in the viral genome, clearance of infection was observed.
Conclusions
This study suggests that severely immunocompromised patients may experience prolonged SARS-CoV-2 infection with insignificant symptoms and persistent virus replication. The authors emphasized the importance of analyzing viral mutations in this group of patients as there is a high possibility of the emergence of variants. Scientists indicated that more research is required to determine the evolution of SARS-CoV-2 in immunocompromised hosts, especially, with regards to transmissibility and virulence.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Nussenblatt, V. et al. (2021) Year-long COVID-19 infection reveals within-host evolution of SARS-CoV-2 in a patient with B cell depletion. medRxiv 2021.10.02.21264267; doi: https://doi.org/10.1101/2021.10.02.21264267, https://www.medrxiv.org/content/10.1101/2021.10.02.21264267v2
- Peer reviewed and published scientific report.
Nussenblatt, Veronique, Allison E Roder, Sanchita Das, Emmie de Wit, Jung-Ho Youn, Stephanie Banakis, Alexandra Mushegian, et al. 2021. “Yearlong COVID-19 Infection Reveals Within-Host Evolution of SARS-CoV-2 in a Patient with B-Cell Depletion.” The Journal of Infectious Diseases 225 (7): 1118–23. https://doi.org/10.1093/infdis/jiab622. https://academic.oup.com/jid/article/225/7/1118/6481528.