
 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
The novel coronavirus SARS-CoV-2 emerged in Wuhan, China late December 2019 and caused the ongoing coronavirus disease 2019 (COVID-19) pandemic. The SARS-CoV-2 Delta variant is the most virulent variant of concern (VOC) relative to Omicron, Gamma, Beta, and Alpha VOCs and causes severe COVID-19 symptoms and a high incidence of mortality.
The first site of interaction of SARS-CoV-2 with human cells is the RBD and is the immunodominant form in the SARS-CoV-2 spike (S) protein. SARS-CoV-2 enters the cell via the interaction between its S and the host angiotensin-converting enzyme (ACE2) receptors. The RBD of Delta contains two new mutations at the T478K and L452R residues.
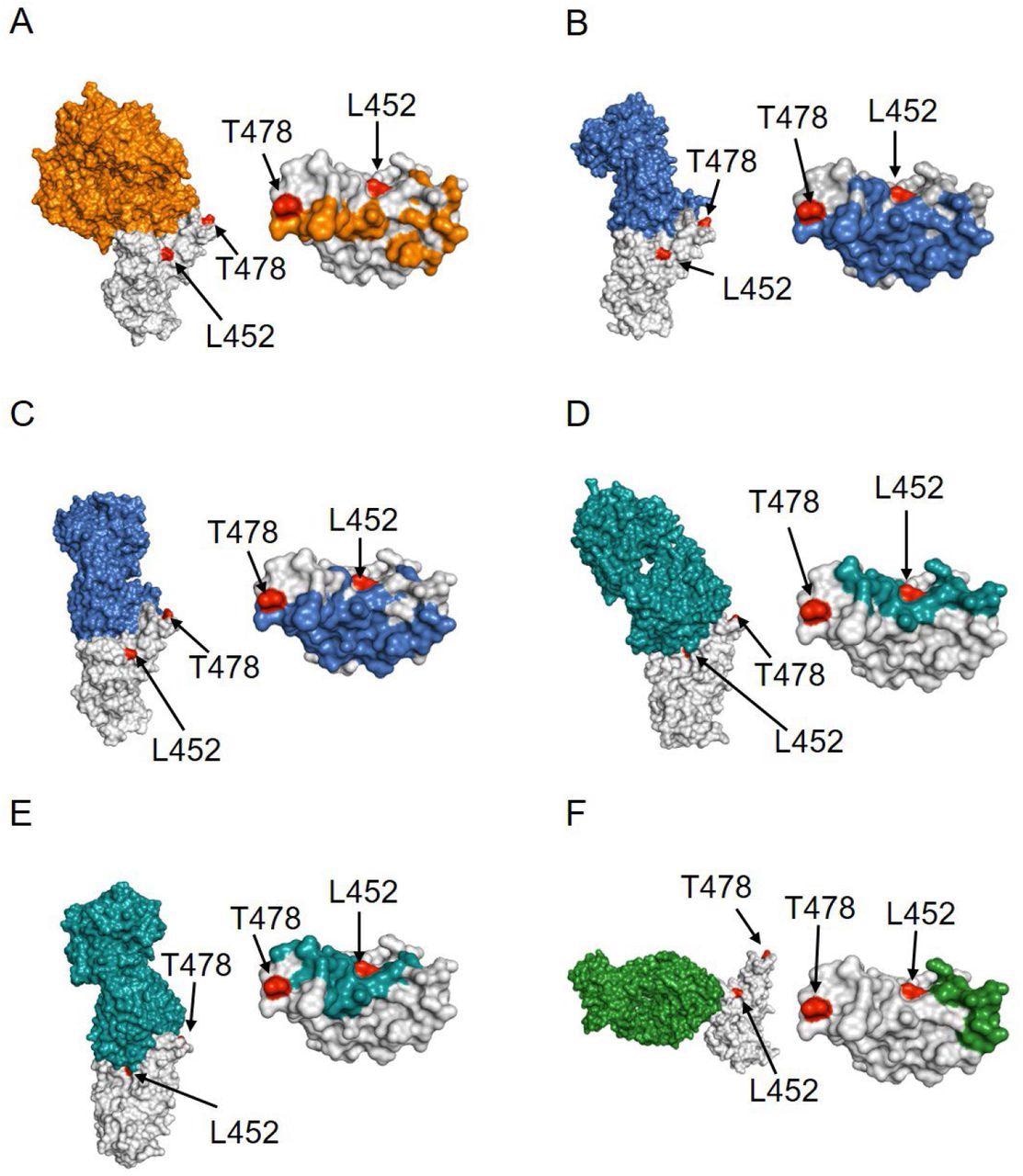
Structural analysis of the location of Delta mutants L452 and T478 (red colored) with respect to RBD (gray colored) complexes with (A) ACE2 receptor (orange colored; PDB ID 6moj), (B) Class 1 antibody CC12.1 (blue colored; PDB ID 6xc2), (C) FDA-approved Class I therapeutic antibody LY-CoV016 (blue colored; PDB ID 7c01), (D) Class 2 antibody P2B-2F6 (teal colored; PDB ID: 7bwj), (E) FDA-approved Class 2 antibody LY-CoV555 (teal colored; PDB ID: 7kmg) and (F) FDA-approved Class 3 antibody REGN10987 (green colored; PDB ID 6xdg). Left panels show the complex structures and right panels show the location of the interacting residues in RBD.
Mutations of previous SARS-CoV-2 VOCs were predicted by in vitro experiments employing parameters like ACE2 receptor binding before their emergence. By contrast, the two novel mutations observed in Delta RBD were not part of the previously discovered variants, and studies have failed to predict their lethality. In addition, biophysical analyses of all SARS-CoV-2 VOCs except Delta such as Gamma, Beta, and Alpha have been conducted. These analyses impart an in-depth understanding of the SARS-CoV-2 variants' host receptor binding, interaction with neutralizing antibodies (NAb), structure, and stability.
About the study
In the current study, the scientists probed the impact of the T478K and L452R mutations of Delta both individually and combined on 1) SARS-CoV-2 RBD expression in human Expi293 cells, 2) RBD stability using thermal and urea denaturation experiments, 3) RBD binding to human ACE2 (hACE2) receptor, and 4) NAbs employing isothermal titration calorimetry (ITC).
Sequences for hACE2 and SARS-CoV-2 RBD were procured from the Uniprot and NAbs from Research Collaboratory for Structural Bioinformatics (RCSB) Protein Data Bank (PDB).
Findings
The results show the RBD of the Delta variant has substantially higher expression relative to the SARS-CoV-2 wild-type (WT) RBD, which was due to the L452R mutation. Despite the Delta mutation's non-conservative character, none of the mutations altered the structure or stability of the RBD.
The binding affinity towards the ACE2 was comparable between the WT and the two Delta mutations. This observation indicates the mutations in Delta did not substantially change the binding affinity of its RBD with the human ACE2 receptor because none of the mutations in Delta directly interacted with ACE2, and it lacks the N501Y mutation, which was associated with the enhanced ACE2 binding.
The mutations in Delta did not allow immune escape from Class one antibodies LY-CoV016 and CC12.1, probably because the two mutations, T478K and L452R, were far away from the binding interface of these antibodies. However, the Delta double mutant L452R/T478K demonstrated no binding towards the Class two antibodies LY-CoV555 and P2B-2F6, as L452R mutation stabilizes the interaction of Delta with these antibodies by forming hydrophobic clusters. Similarly, Delta has a 100-time lower binding affinity towards a Class three antibody REGN10987, contributed by the L452R mutation.
In contrast, since the T478K mutation of Delta lie distant from the binding interfaces of Class two and three antibodies, it did not demonstrate any role in stabilizing inter-molecular interactions and hence was not responsible for the immune escape potential of Delta.
Conclusions
The study findings show that the key biophysical parameter contributing to the fitness landscape of the Delta variant RBD was the L452R mutation-induced immune escape from NAbs instead of the ACE2 binding. Delta did not exhibit increased ACE2 binding and was similar to the SARS-CoV-2 WT variant.
The improved protein expression and immune escape demonstrated by the SARS-CoV-2 Delta variant warrants the significance of robust therapeutic interventions such as advanced monoclonal antibodies (mAbs) in addition to higher vaccination rates to fight Delta and the emergence of future VOCs.
Further, additional therapeutic options are needed to curb the COVID-19 pandemic because of the possibility that Delta might be under continuous evolution and probably result in combinations of mutations that grant immune escape to SARS-CoV-2 from all classes of existing NAbs.
In terms of the currently dominant Omicron variant, it has N501Y mutation and is associated with high ACE2 binding. Although Omicron does not have L452R mutation, it still escapes from the Class two and three NAbs and is likely linked with another set of amino acid mutations. Moreover, if the highly transmissible Omicron combines with the highly virulent Delta, resulting in Deltacrone, it would become a highly lethal SARS-CoV-2 variant.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Biophysical fitness landscape of the SARS-CoV-2 Delta variant receptor binding domain, Casey Patrick, Vaibhav Upadhyay, Alexandra Lucas, Krishna Mallela, bioRxiv 2022.02.21.481311; doi: https://doi.org/10.1101/2022.02.21.481311, https://www.biorxiv.org/content/10.1101/2022.02.21.481311v1
- Peer reviewed and published scientific report.
Patrick, Casey, Vaibhav Upadhyay, Alexandra Lucas, and Krishna M.G. Mallela. 2022. “Biophysical Fitness Landscape of the SARS-CoV-2 Delta Variant Receptor Binding Domain.” Journal of Molecular Biology 434 (13): 167622. https://doi.org/10.1016/j.jmb.2022.167622. https://www.sciencedirect.com/science/article/pii/S0022283622002029.