Dec 6 2013
New insights may explain difficulty of finding drugs for infectious disease
For what is believed to be the first time, researchers at The Johns Hopkins University have illuminated the inner workings of an important class of enzymes located inside the outer envelopes of cells. Much to their surprise, they report, these protein cutters, called rhomboid proteases, are entirely different than nearly every other type of enzyme studied, showing no attraction to the proteins they cut and being extremely slow in making their cuts.
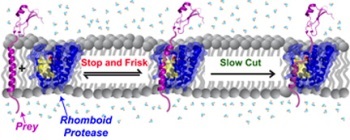
Rhomboid proteases 'stop and frisk' all protein 'prey' in the membrane indiscriminately. The rhomboid protease cuts slowly so that stable proteins can escape from being cut, but too slow unstable proteins get the chop.
The researchers say their findings may explain why efforts to design drugs targeting the enzymes, which are involved in Parkinson’s disease and parasite infections, have so far failed. A summary of their research will be published in the journal Cell on Dec. 5.
“The properties that would be seen as a problem for most enzymes are precisely what give rhomboid proteases their unique function in the cell,” says Sinisa Urban, Ph.D., associate professor of molecular biology and genetics at the school of medicine and the Howard Hughes Medical Institute. “Our results indicate that to catch and cut all unstable proteins in cell membranes, rhomboid proteases must ‘stop and frisk’ all of them indiscriminately, and their cut rate must be slow enough to allow stable proteins to escape without injury.”
Enzymes are essentially proteins that modify molecules, such as other proteins. In comparing enzymes, scientists analyze their efficiency, which depends on two factors: the “magnetism” or attraction between an enzyme and its “prey” and the speed with which the enzyme makes its modifications.
Most enzymes work in a watery environment, which makes them relatively easy to study in laboratories. However, the oily layer of fat molecules that creates a boundary between the inside and outside of a cell is a drastically different environment. It is thicker; it lacks water, which is usually required for enzymes to work; and it is two-dimensional, which limits the positioning of enzymes and their prey. Despite these differences, many researchers had assumed that all enzymes worked in the same way, whether they resided in watery or oily environments.
To explore that assumption, Urban's team needed to test the speed of rhomboid proteases. But Seth Dickey, Ph.D., the lead author of the study, explains that measuring the speed of a membrane enzyme the traditional way was something like having people begin a race inside a mile-long tunnel where it can’t be seen. “We didn’t have the tools to allow us to see the beginning of the race, when the enzymes are fastest and just starting their work,” he says.
To overcome this difficulty, Urban’s team first synchronized the enzymes by setting up their “race” under acidic conditions, which inactivates rhomboid proteases. In this way, the researchers were able to turn the enzyme off and keep it off until they neutralized the acidity when they were ready to begin monitoring the race.
What the team witnessed were among the slowest enzymes ever, Urban says. They took more than two and a half minutes to make each cut, whereas most enzymes take only hundredths of a second. In addition to being slow, the enzymes showed no attraction to some proteins over others, as most enzymes do.
While these findings were surprising, Urban thinks there is a reasonable explanation. Normally, rhomboid proteases are probably acting as quality control enzymes that cut up unstable proteins before they can do any damage to the organism, he suggests. In watery environments, unstable proteins are quite wobbly and are easily recognized and destroyed by other quality control enzymes. Recognizing unstable proteins in oily environments is more difficult because fat molecules support the proteins and give them the appearance of being stable.
From their previous work, the team knows that rhomboid proteases resemble water-filled barrels with side gates for protein entry. Their latest work suggests that stable and unstable proteins alike enter the side gate. Stable proteins likely remain intact and drift back out into the membrane before the enzyme’s slow scissors have a chance to clip them, while unstable proteins start to wobble in the watery interior and have a hard time exiting the enzyme’s barrel. There, the enzyme will clip them, allowing the cell to recycle them for spare parts.
“We hope this new enzyme synchronizing technique and the insights we made will be widely applied to all membrane-embedded enzymes,” says Urban.
He notes that membrane-embedded enzymes include enzymes that have been implicated in a wide variety of diseases: from Alzheimer’s disease to leukemia to viral, bacterial and fungal infections. For example, gamma secretase is a membrane enzyme in charge of processing the protein beta-amyloid. Clipping beta-amyloid in the wrong place seems to lead to clumps of the protein in the brains of Alzheimer’s patients. Urban says that a proper understanding of membrane enzymes is paramount to finding drugs that modify their behavior.