The current COVID-19 pandemic demonstrates the vast unknown of virology, which continues to challenge the ability of humanity to remain healthy when faced with pathogens. While most known microbes have restricted affinity for specific species, continuing to adapt with the host species, the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has crossed over from an unknown animal reservoir, like the preceding SARS and MERS coronaviruses, to infect human cells. Such viruses are typically more readily infective and cause more severe disease, as they have not yet adapted fully to the target host.
.jpg)
Novel Coronavirus SARS-CoV-2 Colorized scanning electron micrograph of an apoptotic cell (green) heavily infected with SARS-COV-2 virus particles (purple), isolated from a patient sample. Image captured at the NIAID Integrated Research Facility (IRF) in Fort Detrick, Maryland. Credit: NIAID
The burning question is how novel viruses acquire the ability to recognize, bind to and enter human cells for the first time – whether this is dependent only on viral proteins recognizing host cell proteins, or adaptations in other viral processes that allow replication in a human host.
This issue is discussed by researchers at the University of Calgary in a new study published on the preprint server bioRxiv* in June 2020. The spike protein is the most well known of the SARS-CoV-2 proteins, and its binding to ACE2 receptors on the host cell is responsible for viral entry into the target cell. The human ACE2 (hACE2) has some rare variants which make the host more vulnerable to infection. Similarly, the spike protein of this virus has a greater affinity for the receptor than the previous SARS virus, which is another possible explanation for the increased infective potential of the current virus.
The Study: Origin of ACE2 Binding Affinity
The current study examines the origin of this spike protein variant with its affinity for hACE2, using molecular dynamics (MD) simulations along with sequence reconstruction to identify the adaptation pathway of the virus. The result is a preliminary phylogenetic analysis that agrees with earlier studies – the virus is 96% similar to the bat coronavirus (RaTG13) genome and 90% similar to the Pangolin-CoV genome.
The next step was to carry out a more detailed analysis of 479 sequences collected from December 30, 2019, to March 20, 2020, where they found 16 variants. Of these, 11 were missense mutations occurring in 5% or more of cases, and each had its own phylogenetic route.
The researchers then tried to recreate the ancestral sequence for the spike-RBD region, so that they could identify the important mutations that specifically drive its recent adaptation to the human host. They reconstructed the hypothetical common ancestor spike-RBD sequence for all human SARS-CoV-2 cases, called N1, and for the common ancestor with the closest animal virus, called N2.
N1 is identical to the sequence in the SARS-CoV-2 reference sequence, but the N0 sequence is unique, which shows that this virus has originated uniquely. The two differ at 4 positions. The ancestral protein gave rise to various descendants, one of which is the RaTG13. Since this was around in 2013, the researchers conclude that the ancestral strain existed as early as that year, at least. In other words, the N0-N1 branch has been evolving for at least 7 years.
Ancestral Sequence Had Higher Binding Affinity
What are the functional differences between N0 and current spike-RBD sequences? The researchers used MD simulations of the spike-RBD-hACE2 complex, beginning with the X-ray crystal structures. The model showed that the free binding energy for this complex decreased as N0 changed to N1. Thus, this actually reduced the binding affinity both in the simulations and in vitro.
However, two of the changes were associated with more significant decreases than the other. This shows that the N0 strain had, unexpectedly, greater binding affinity than the N1 strain. This is the first study to show that the common ancestor of both SARS-CoV-2 and the RaTG13 had the ability to bind to the ACE2 receptor in humans.
Other Molecular Changes Key to Infectivity
The implications are that firstly, the binding affinity of the spike-RBD to hACE2 is not the primary driver of the highly infectious nature of the current virus since the ancestral virus was capable of doing this too. Secondly, the researchers suggest that this virus was, even then, able to bind tightly to the receptor. Therefore, this was not sufficient to produce the currently observed ability to spread rapidly and widely among humans. Instead, this must be due to another set of mutations in the viral genome.
Yet another implication is that the current virus may not have jumped to humans from an animal origin at all because its affinity for hACE2 was not a recently acquired molecular trait. This may mean that the ability to infect human cells was present over a more extended period in the past, but produced less obvious or fewer clinical symptoms which passed unnoticed. Another alternative was that it affected only a small number of people, allowing it to remain under the public health radar.
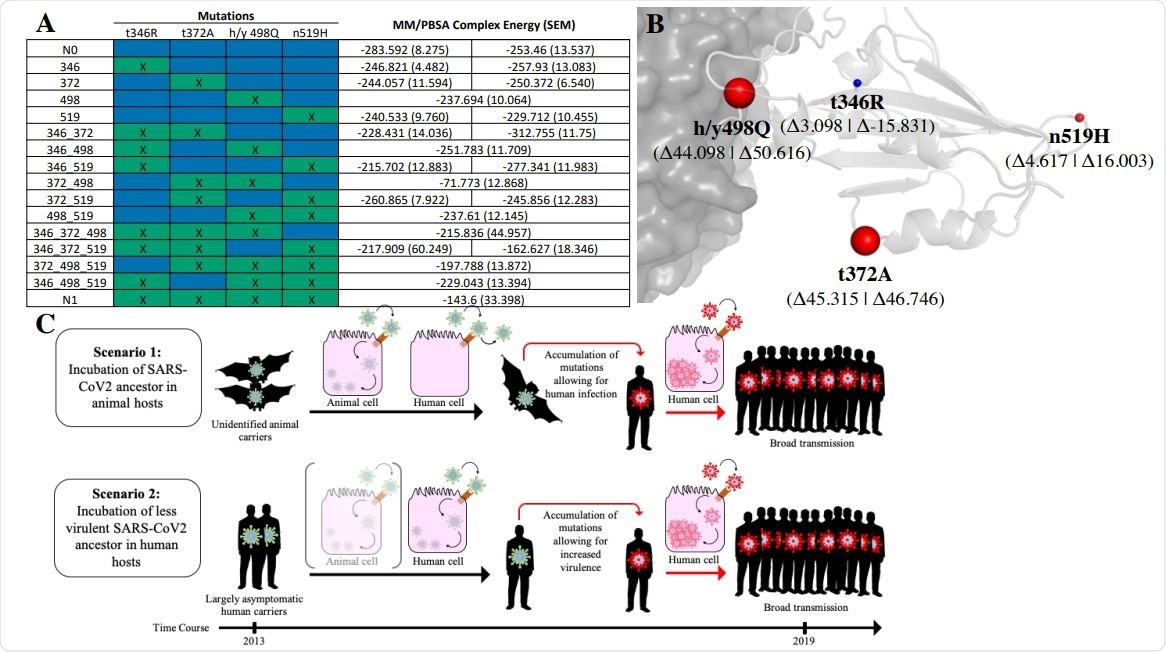
Characterization of SARS-CoV-2 Spike-RBD functional evolution. A. Table of MM/PBSA binding energies between receptor binding domains of SARS-CoV2 evolutionary constructs and hACE2 receptor (note that lower energy indicates tighter binding). Blue cells indicate the presence of the ancestral (N0) state and green cells (with an “x”) indicate the presence of the SARS-CoV-2 state (N1) at a given position. Two values are present for constructs with an ancestral (N0) state at position 498 (which reflect the ambiguity of its ancestral reconstruction), corresponding to h498 and y498 from left to right. Energies are shown as the mean of three replicate simulations with SEM indicated in parenthesis. B. The relative effect of changes in the SARS-CoV-2 receptor-binding domain from ancestral (N0) to SARS-CoV-2 (N1) state on MM/PBSA binding energies. Size of spheres indicates the relative magnitude, with red spheres indicating decreased binding affinity and blue indicating increased binding affinity. Values are averaged for h498 and y498 states (both raw values shown in parentheses). C. Schematic of two possible evolutionary scenarios stemming from the observed evolutionary SARS-CoV-2 Spike-RBD function. In Scenario 1, it is postulated that a zoonotic ancestral SARS-CoV-2 strain possessed the ability to effectively bind hACE2 but was unable to effectively enter human cells, requiring the presence of subsequent mutations to infect humans. In Scenario 2, an ancestral SARS-CoV-2 strain was actively infecting humans prior to the outbreak at low levels, thus escaping public health detection until subsequent mutations lead to increased infectivity and/or severity.
Future Research
These possibilities can only be tested by a broad-spectrum approach to sequencing all coronavirus strains in human populations, as this will reveal the presence of closely related viruses if such are present.
The current study is an in silico study, and further validation of these findings is necessary using combinatorial libraries which can be screened to map functionalities to the genomic regions of the virus. This will help understand how the virus evolved in the most recent past.
The researchers conclude: “It appears that the SARS-CoV-2 Spike-RBD did not recently evolve binding affinity to a human-specific protein. Instead, that function appears to have been latent, making it clear that the evolution of this disease – along with so many other aspects of its etiology – is more complex than expected.”

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources