As the COVID-19 pandemic continues to cause economic and social disruption in addition to infections and deaths, it is slowly becoming clear that the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is changing as it spreads to different regions. This trend is driven by spontaneous adaptive mutations, shaped by the different selection pressures exerted by varying environmental conditions and hosts. This has been enhanced by the adoption of lockdowns and other social distancing measures. Now, a new study published on the preprint server bioRxiv* in August 2020 shows the importance of uncovering the dynamics of the virus with particular attention to geographic variation.
The current study was focused on finding and interpreting the association between the evolution of the SARS-CoV-2 and the conditions in the Indian environment and hosts.
Increasing Spike Mutations, Lower Viral Fitness
The researchers looked at how the viral spike glycoprotein showed mutation-derived differences since this is the key to viral binding, fusion, and cell entry leading to host cell infection. This is the key pathogenetic feature that differentiates it from earlier pathogenic coronaviruses SARS-CoV and MERS-CoV. It also carries the two most important genetic features of the virus.
As a result, the researchers say, “By examining the architecture of the spike glycoprotein and its mechanics, one can reveal the susceptibility of the virus and can foresee the facts helping in the discovery of the SARS-CoV-2 antidote.”
They used the genomic sequences of 630 Indian isolates retrieved from the GISAID database. Tracing these mutations showed that the spike protein variants followed two pathways starting from two ancestral strains, namely, Wuhan-Hu-1/2019 and its D614G variant.
While the Wuhan-Hu-1/2019 strain gave rise to the D614G variant, it also evolved to 20 other variants, of which one further led to three others. The other ancestral strain D614G gave rise to 47 variants, of which 9 led to further variants. The two ancestral strains are predominant, but there were 16 detectable spike protein variants found in multiple isolates, with different levels of spike-receptor stability. Strikingly, several mutations occurred independently at the same position across eight variants, which were not linked phylogenetically.
Over half of the variants were found only in India, but over two-thirds had lower spike-receptor stability than the ancestral strains. This should be expected in a region where a new virus that is rapidly mutating tries to adapt to a host while doing so. Both positive and detrimental mutations may occur without reversion in the absence of any genetic recombination mechanism.
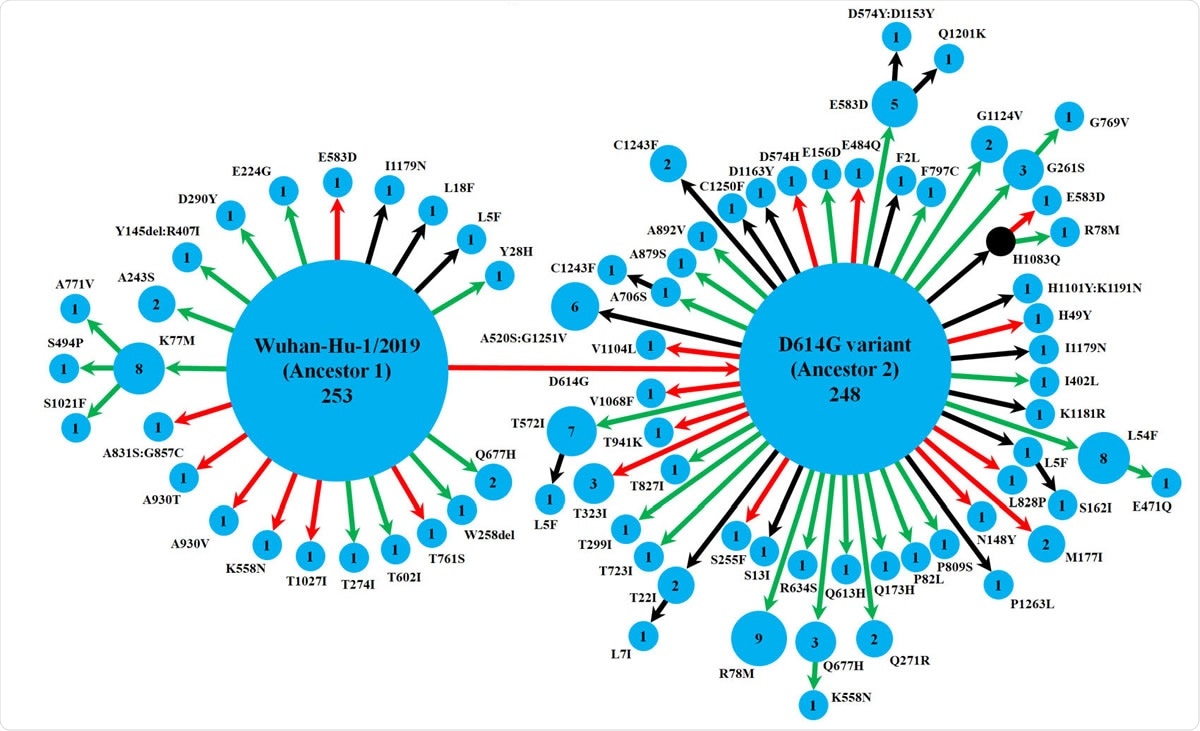
Schematic representation of the diversity of spike protein variants circulating in India, using maximum likelihood-based phylogeny reconstruction. Each node represents a specific spike protein variant, while the node-size and the number inside depict the frequency of that variant. The red or green color of each arrow indicates the higher or lower stability index respectively of the S-R complex for each variant than the major ancestral variant it emerged from (either Ancestor 1 or Ancestor 2). The black arrows lead to the variants for which the docking scores could not be determined either because of the presence of at least one variation outside the available template region for docking(18, 42), or due to non-existing isolate in the lone hypothetical node with H1083Q mutation denoted by black color. This black node signifies a variant with no available isolate in our dataset, while it gives rise to two derived variants, H1083Q:R78M and H1083Q:E583D, for which representative isolates were available.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
The implication is that this rapid buildup of mutations in the more recent strains leads to a loss of fitness compared to their ancestors, despite the constant increase in the rate of mutations. The researchers invoked Muller’s ratchet, the hypothesis that in asexual reproduction, the irreversible accumulation of harmful mutations causes a species to die out gradually.
The interesting thing is that scientists perceive the ability of SARS-CoV-2 to jump the species barrier and infect humans to be due to specific mutations. This is an RNA virus which lends it an inherently high mutation rate. Understanding how this is occurring at the molecular level could help understand how it causes disease and how it can be curbed.
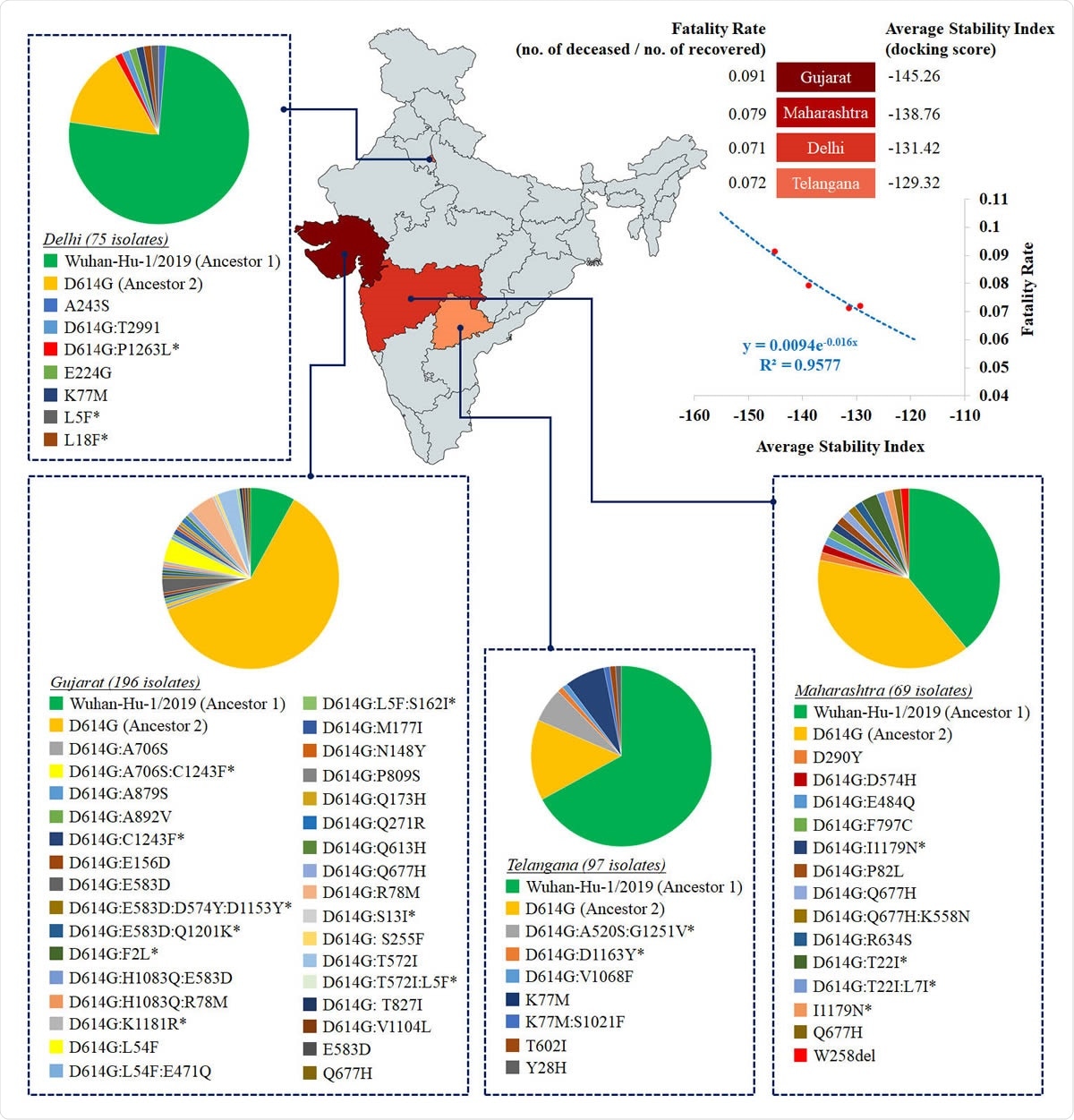
Heat map distribution across four Indian states with >50 sequenced isolates based on average stability index. The average stability index for a particular state denotes the averaged value of docking scores / HADDOCK scores of S-R complexes for all circulating variants. The values of average stability index and fatality rate in Indian states are plotted to fit an exponential function (R2=0.96).
Lower Fatality Rate with Decreasing Stability
The most significant spike protein diversity was in Maharashtra, Odisha, West Bengal, and Gujarat, but this is limited by the fact that many other states had very few isolates. This diversity could be due to positive selection pressures enabling the protein, which is a prime immune target, to evade host immunity and allow entry into the cell. This could affect spike-receptor binding. They found 41 and 22 mutations in the S1 and S2 subunits of the spike, respectively.
The study of mean spike-receptor complex stability in each state, based on the individual stability indices of the variants in circulation in that state, was then performed using the docking score of the S1 subunit with the receptor. Limiting it to only those with 50 or more sequenced isolates, they looked at this index in Maharashtra, Delhi, Gujarat, and Telangana, which account for 70% of the total isolates, and cover a wide range of diversity.
The mean stability of the spike-receptor complex across these variants was strongly and exponentially correlated with fatality rates (deceased: recovered cases) across the Indian subcontinent. The fatality rate dropped by seven times between April 11 and June 28, 2020. Telangana and Delhi had similar average stabilities at a 7% fatality rate, while Maharashtra and Gujarat showed exponentially lower stabilities, with 8% and 9% fatality rates, respectively.
They say that this suggests, “the S-R complex stability [is] a potential marker to assess the severity of the disease.” They suggest that further study is necessary at the population level to validate this experimentally, especially since the actual parameter (the docking score) used has not been directly correlated with the fatality rate so far.
Implications
The researchers put forth the theory that this loss of stability may be responsible for the lower national fatality rate. In India, after the initial phase in which the fatality steadily went up for four weeks, to peak at 38% on April 11, 2020, it declined sharply until it reached 5% on June 28, 2020, the date at which the researchers last obtained their data. However, the mutation rate was increasing during this period from March to May 2020.
They suggest, “Could Muller’s ratchet be a player in shaping SARS-CoV-2 evolutionary dynamics in India?” In other words, the virus could be gradually extinguishing itself under the accumulating weight of deleterious mutations, without the washout process of natural selection being able to compensate because of the rapid rate of mutations. This causes harmful mutations to be fixed in the viral population, and this buildup could lead to a “mutational meltdown.”
The short period of the study precludes a definitive answer, but the trend may be seen, according to the researchers. This could also help evolve a better therapeutic approach, facilitating the meltdown process by inducing more harmful mutations. This will require large-scale genomic studies to answer the question, and thereby helping to improve public health surveillance and prevention programs.
The researchers sum up: “Altogether, while we propose the potential of SR complex stability to track disease severity, we urge an immediate need to explore if SARS CoV 2 is approaching mutational meltdown in India.’

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources