Researchers from the Vaccine and Infectious Disease Division at the Fred Hutchinson Cancer Research Center, USA, have revealed that adaptive evolution in the antigenic regions facilitates seasonal coronaviruses to escape the host immune responses and to cause recurrent infections.
Since their identification in the 1960s, human coronaviruses - including OC43, 229E, HKU1, and NL63 - are known to be responsible for at least 15% of common colds. Seasonal infections caused by these viruses generally peak in January – March in the Northern Hemisphere. In the general human population, these viruses cause mild respiratory infections; however, in aged people or in people with weakened immune systems, severe infections may occur.
Three new human coronaviruses have been identified in recent years, which are known to cause severe respiratory infections in humans. These viruses include severe acute respiratory syndrome coronavirus (SARS-CoV-1), Middle-East respiratory syndrome coronavirus (MERS-CoV), and SARS-CoV-2 (the causative pathogen for coronavirus disease 2019; COVID-19).
To cause repeated infections in humans, a virus must acquire genetic mutations that cause adaptive alteration in the antigenic viral region. This allows the virus to escape the humoral immune responses of the host. Some seasonal human respiratory viruses, such as the influenza virus, are known to undergo antigenic evolution. Vaccines developed against such viruses thus need to be reformulated frequently to tackle viral spread.
In the case of human coronaviruses, the most antigenic protein is the spike protein, which is a viral surface protein responsible for viral entry into host cells. Studies have shown that the predominance of certain genetic variants of human coronaviruses is associated with positive selection in the viral spike protein.
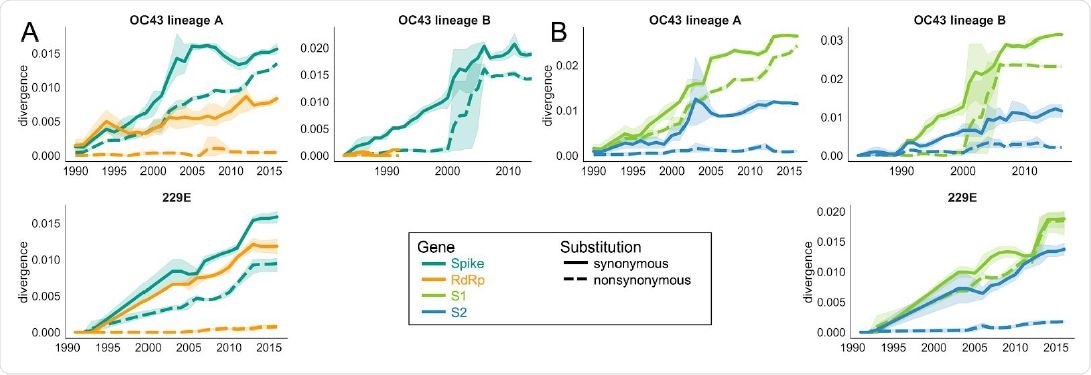
Nonsynonymous divergence is higher in OC43 and 229E Spike S1 versus S2 or RdRp. A: Nonsynonymous (dashed lines) and synonymous divergence (solid lines) of the spike (teal) and RdRp (orange) genes of all 229E and OC43 lineages over time. Divergence is the average Hamming distance from the ancestral sequence, computed in sliding 3-year windows which contain at least 2 sequenced isolates. Shaded region shows 95% confidence intervals. B: Nonsynonymous and synonymous divergence within the S1 (light green) and S2 (blue) domains of spike. Year is shown on the x-axis. Note that x- and y-axis scales are not shared between plots.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Current study design
The scientists used computational methods to analyze adaptive evolution in the spike protein and RNA-dependent RNA polymerase of four seasonal coronaviruses (OC43, 229E, HKU1, and NL63) that are known to cause repeated infections in humans. Specifically, they analyzed the mutations that change the amino acid sequence of a protein (nonsynonymous substitution). Additionally, they estimated the rate of adaptive substitutions and the Time to Most Recent Ancestor (TMRCA). TMRCA is the timescale of population turnover used to quantify the positive selection.
Important observations
The scientists used publicly available genetic sequences of human coronaviruses to create a phylogenetic tree. Through phylogenetic analysis - the study of the evolutionary development of a species or a group of organisms or a particular characteristic of an organism - they observed that several co-evolving lineages are present for OC43. In OC43 and 229E lineages, the average number of mutations at each position was found to be higher in S1 subunit of the spike protein compared to that in S2 subunit.
Regarding the rate of mutations, they observed a significantly higher rate of nonsynonymous mutation in the spike protein of OC43 and 229E compared to that in the RNA-dependent RNA polymerase. Moreover, the nonsynonymous divergence increased progressively in the spike protein; whereas in RNA-dependent RNA polymerase, it remained constant. These findings indicate positive selection on the spike protein and purifying selection on the RNA-dependent RNA polymerase.
Regarding the rate of adaptive substitution, they observed that the S1 subunit of the OC43 gathers 0.45 – 0.56 adaptive substitutions each year. Similarly, the S1 subunit of 229E spike protein gathers 0.26 adaptive substitutions each year. They further observed that the rate of accumulation of adaptive substitutions in the influenza virus's receptor-binding domain is 3 times faster than that in OC43 and 229E. Taken together, these findings indicate that the S1 subunit of the OC43 and 229E spike protein is under positive selection, which can give rise to new genetic variants.
By estimating and analyzing the TMRCA values, the researchers observed robust directional selection in the S1 subunit of both OC43 and 229E, which may be driven by pressures to escape host immune responses.
The take-home message
The S1 subunit of the spike protein of human coronaviruses (OC43 and 229E) is under adaptive evolution. The rate of accumulation of adaptive mutations is about one-third of the influenza virus rate. The antigenic evolution observed in S1 may be selectively favoring viruses to escape host immune responses. However, no evidence of adaptive evolution is observed for other seasonal coronaviruses, such as NL63 or HKU1.
Given the genetic similarity between OC43 and SARS-CoV-2, it might be expected that SARS-CoV-2 also evolves selectively in S1; and in that case, COVID-19 vaccines that are presently under final stages of investigations may need to be frequently reformulated to effectively eliminate circulating viral variants.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources