Researchers of the Hubei University of Medicine, China, have analyzed several mutations in different variants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) from mammal species that suggest natural selection and host adaptation are occurring in the virus.
SARS-CoV-2 is the causative agent for the coronavirus 2019 (COVID-19) pandemic, which continues to be a global crisis.
While the virus can transmit from human-to-human hosts easily, multiple cases of human-to-animal and animal-to-human transmission have also been observed.
A wide range of animals has been observed or suspected to be susceptible to the virus. Cats, dogs, lions, tigers, gorillas, snow leopards, white-tailed deer, ferrets and mink have all tested positive for having acquired SARS-CoV-2.
Most notably, mink in farms in The Netherlands were reported to be contracting the virus from humans, but then also transmitting the virus both to mink and back to humans.
Thus, keeping the spread of the virus under control is essential to prevent local outbreaks in both wild and captive animal populations. Acknowledging the mutations SARS-CoV-2 variant strains acquire in different animal species is also crucial in achieving this goal.
The study
Professor Long Liu, of the Hubei University of Medicine, and colleagues analyzed multiple variants of SARS-CoV-2 found in different mammal species and compared these to the initial “Wuhan” circulating strain of the virus during the first wave of the pandemic.
The team used SARS-CoV-2 genome sequences from humans, cats, dogs, tigers, lions, ferrets and mink.
These sequences were used to construct a phylogenetic tree to infer the evolutionary history of the viral genomes. The researchers then used an adaptive evolution server (Datamonkey) to identify branches where the virus had been under selective pressure.
Following this, Liu and colleagues used the codon adaptive index (CAI) to assess the variety of mutations between the different variants.
SARS-CoV-2 binds to angiotensin-converting enzyme 2 (ACE2), an enzyme located on the outside of (primarily) respiratory cells, to access the cell. The researchers recreated the structures of human ACE2 and the crystal complex of SARS-CoV-2 to model binding efficiency between variants. Structures of mink ACE2 and the mink viral spike were also constructed and used in the analysis.
Results
The researchers found that SARS-CoV-2 of the mink cluster group had more substitution mutations compared to the reference strain.
Substitutions of cytidine in mink-CoV account for nearly 50% of the substitutions, while in other animals, cytidine accounts for only 30% of the substitutions,” say the authors, “The substitution of adenine in SARS-CoV-2 in other animals is threefold higher than that in mink-CoV.”
Cytidine-to-uracil mutations occurred over 40% of the time and 8x more so than mutations of the reverse. Substitution of adenine and guanine was also 3x higher in nonsynonymous mutations than in synonymous mutations.
Notable and convergent mutations were also observed in the spike protein of the mink SARS-CoV-2 variants. Variations in the spike protein were observed in all variants for animal species from the reference strain, D614G and A222V, most commonly found. The researchers found that these sites, as well as sites 262 and 439, were exposed to strong positive selective pressure, while other sites (294, 413, 1018 and 1100) were negatively selected for.
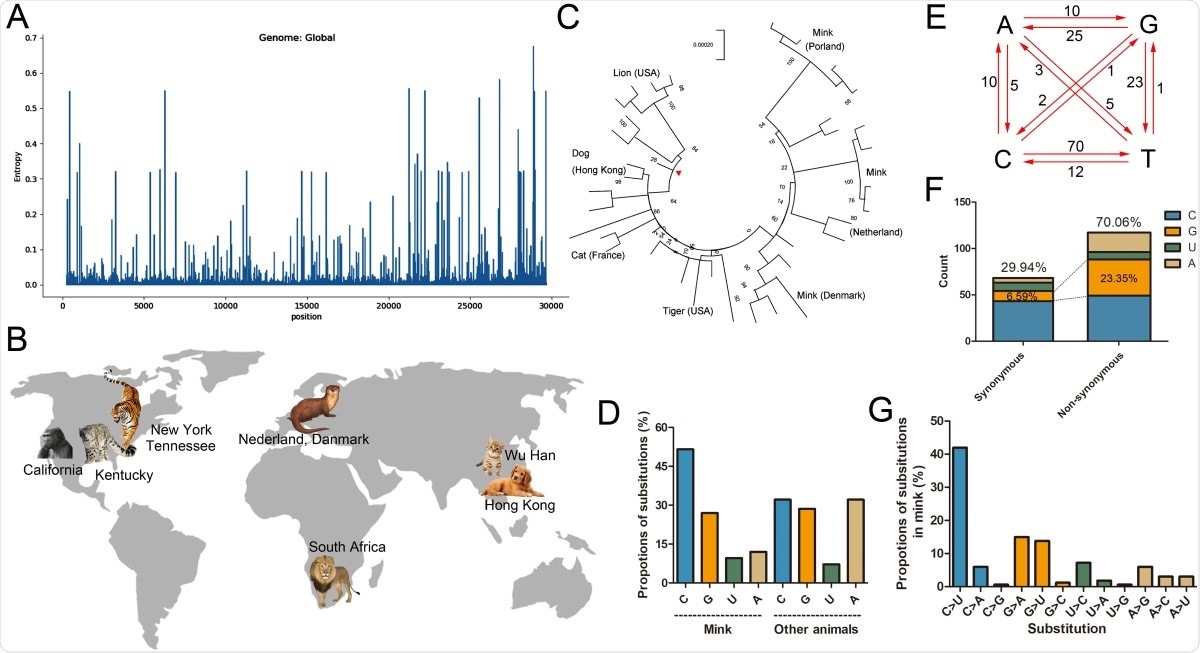
Composition and substitution analysis of SARS-CoV-2 isolated from animals. (A) The evolutionary entropy of specific sites on the SARS-CoV-2 genome in all the GISAID sequences on February 1, 2021. (B) The reported animals infected with SARS-CoV-2 and the defined transmission route from human to animal. (C) Phylogenetic tree using the maximum likelihood method and Tamura-Nei model performed by MEGA-X. The tree was provided with 500 bootstraps. (D) The proportions of uracil, guanine, thymine, and cytidine substitutions (nonsynonymous) in mink SARS-CoV-2 and other animals were separately counted. (E) Base pair changes observed in the mink SARS-CoV-2 genomes. All transitions and transversions were recorded and analyzed (see Supplementary Table S2). (F) The synonymous and nonsynonymous substitutions of mink-CoV were counted and analyzed. (G) The relative proportions of all transitions and transversions were separately analyzed.
To determine the optimization of SARS-CoV-2 to hosts, the researchers calculated the average CAI of the spike region and the full genome of the virus. Higher CAI scores correspond to increased host optimization; these scores were lower (compared to humans) in pangolins, cats, dogs, tigers, and lions, but higher in bat hosts.
Mink SARS-CoV-2 additionally showed greater preference for mink ACE2 over human ACE2, following mutation at site 453 from Y453 (found in humans) to F453.
Concluding remarks
SARS-CoV-2 is suspected to have had a zoonotic origin, from either bat or pangolin coronaviruses. Its willingness to jump intraspecies hosts should be of great concern.
Professor Liu and her team have identified evidence for natural selection and host adaptation in SARS-CoV-2 in mink, with a great number of variations also found between other mammal hosts. Thus, understanding these mutations and the selective pressures that drive them is critical in preventing ecological outbreaks or new variants of the virus from arising.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Lei Z., et al. Substitutions and codon usage in SARS-CoV-2 in mammals indicate natural selection and host adaptation. bioRxiv, 2021. doi: https://doi.org/10.1101/2021.03.15.435309, https://www.biorxiv.org/content/10.1101/2021.04.04.438417v1
- Peer reviewed and published scientific report.
Fu, Yanan, Yanping Huang, Jingjing Rao, Feng Zeng, Ruiping Yang, Huabing Tan, Zhixin Liu, Weixing Du, and Long Liu. 2022. “Host Adaptation of Codon Usage in SARS-CoV-2 from Mammals Indicates Potential Natural Selection and Viral Fitness.” Archives of Virology, September. https://doi.org/10.1007/s00705-022-05612-6. https://link.springer.com/article/10.1007/s00705-022-05612-6.