It is widely assumed that the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) resulted from a zoonotic transition from bats because mutations occurred in the viral spike protein that allowed it to interact with the human ACE2 receptor.
The spike protein of a closely related bat virus named RaTG13, bearing 96% genome sequence similarity with SARS-CoV-2, bears striking structural resemblance with that of SARS-CoV-2, though it does not efficiently interact with the human form of the ACE2 receptor.
In a paper recently uploaded to the preprint server bioRxiv* by Kirchhoff et al. (May 31st, 2021) it is demonstrated that a single mutation to the RaTG13 spike protein results in significantly greater affinity towards the human receptor, indicating that the presence of a positively charged amino group at this position better facilitates interaction with the human ACE2 receptor, and helping to identify other viruses with zoonotic potential.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Spike protein mutations
Currently recognized differences in structure between the spike proteins of SARS-CoV-2 and RaTG13 include specific alterations to the receptor-binding domain and a four amino acid insertion that comprises a furin cleavage site, a critical factor in the extreme pathogenicity of COVID-19. However, it is not yet known how the bat virus from which SARS-CoV-2 mutated may have initially crossed the species barrier, as some specific features must have emerged beforehand to allow interaction with the human ACE2 receptor. The group identified amino acid R403 of the spike protein as involved in intramolecular interactions that stabilize the active form and contribute significantly to interactions with ACE2. Comparative studies of genome databanks revealed that this amino acid is very highly conserved, with only 233 records indicating a conservative R403K mutation and an additional 18 demonstrating deletion, out of a total of 1.7 million SARS-CoV-2 sequences.
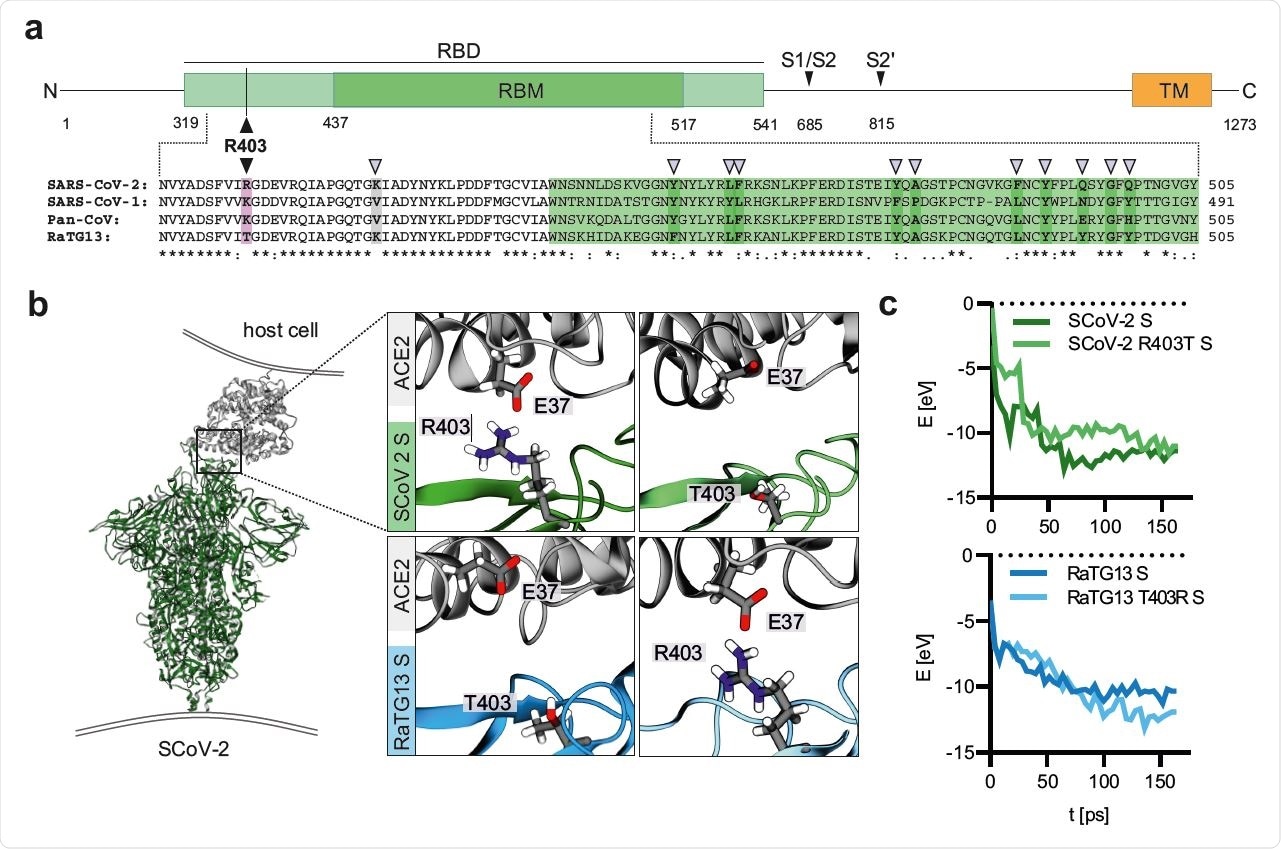
Modeling of the interaction of Coronavirus Spike residue 403 with human ACE2. a, Schematic representation of the SARS-CoV-2 S protein (top panel), domains are indicated in different colors. Receptor binding domain (RBD), light green. Receptor binding motif (RBM), dark green. Transmembrane domain (TM), orange. R403, pink. S1/S2 and S2’ cleavage sites are indicated. Sequence alignment of SARS-CoV-2, SARS-CoV-1, Pan-CoV and RaTG13 Spike RBD (bottom panel). Sequence conservation is indicated. purple arrows denote important residues for ACE2 binding. b, Reactive force field simulation of SARS-CoV-2 Spike in complex with human ACE2 (PDB: 7KNB) (left panel) and focus on position 403 in SARS-CoV-2 S (R) or RaTG13 S (T) or respective exchange mutants at position 403 (right panel). c, Exemplary energy curve of the reactive molecular dynamics simulation for SARS-CoV-2 S and SARS-CoV-2 S R403T (top panel) and RaTG13 and RaTG13 T430R spike with human ACE2 (bottom panel)
Both SARS-CoV-2 and SARS-CoV-1, also emerging from bats, bear a positively charged residue at position 403, while the bat virus RaTG13 has a neutral threonine amino acid at this position. Docking simulations indicate that this mutation produces far greater affinity towards the receptor. The group further demonstrates this in vitro using modified viral vectors expressing the neutral amino acid 403 spike protein, which entered Caco-2 cells 40% less efficiently than those bearing a positively charged residue at this position.
The RDG tripeptide motif (Arginine-Glycine-Aspartic Acid) is associated with cell membrane adhesion via interaction with transmembrane integrin proteins. It has been suggested that the R403 residue of SARS-CoV-2 may produce such a binding site that aids in cell entry and contributes to the high infectiousness of the virus. To test this theory, the group applied integrin inhibitors to both SARS-CoV-2 and RaTG13 during infection, noting no significant difference and concluding that the enhanced affinity of SARS-CoV-2 towards ACE2 over RaTG13 is likely due to the demonstrated enhanced interaction with the receptor, as opposed to by integrin interaction.
Intestinal organoids were also utilized to test the ability of each virus to infect the tissue while bearing particular residues at position 403. SARS-CoV-2 bearing the usual R403 amino acid performed as expected, infecting the organoids through interaction with ACE2, while the R403T mutant showed attenuated effects. The RaTG13 virus bearing the T403 amino acid was unable to infect the organoids while introducing the T403R mutation allowed significantly greater infection of the human cells. Similar results were observed in a wide range of cells that were engineered to express the human ACE2 receptor, with the SARS-CoV-2 version of the spike protein bearing a positive residue at position 403 better infecting all cells.
Interestingly, when the group applied the wildtype SARS-CoV-2 spike protein (R403) or the mutated RaTG13 spike protein (T403R) to cells heavily overexpressing the bat ACE2 receptor, low to moderate levels of infection were observed. However, the wildtype RaTG13 spike protein (T403) is completely unable to infect these cells, suggesting that RaTG13 uses an alternate route of infection other than the ACE2 receptor in bats while still being capable of entry via this route when the receptor is overexpressed.
A number of other key amino acids are involved in the interaction between the SARS-CoV-2 spike protein and the ACE2 receptor, which explains why cell entry is merely impeded rather than prevented in the absence of the positively charged residue at position 403.
The group state that the majority of other RaTG13-related bat coronaviruses already bear a corresponding positively charged residue at this position, suggesting that whichever bat virus SARS-CoV-2 actually emerged from likely already bore this mutation, which allowed initial zoonotic transition and the acquisition of other mutations that enhanced affinity towards the human ACE2 receptor.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Spike mutation T403R allows bat coronavirus RaTG13 to use human ACE2, Fabian Zech, Daniel Schniertshauer, Christoph Jung, Alexandra Herrmann, Qinya Xie, Rayhane Nchioua, Caterina Prelli Bozzo, Meta Volcic, Lennart Koepke, Jana Krüger, Sandra Heller, Alexander Kleger, Timo Jacob, Karl-Klaus Conzelmann, Armin Ensser, Konstantin M.J. Sparrer, Frank Kirchhoff, bioRxiv, 2021.05.31.446386; doi: https://doi.org/10.1101/2021.05.31.446386, https://www.biorxiv.org/content/10.1101/2021.05.31.446386v1
- Peer reviewed and published scientific report.
Zech, Fabian, Daniel Schniertshauer, Christoph Jung, Alexandra Herrmann, Arne Cordsmeier, Qinya Xie, Rayhane Nchioua, et al. 2021. “Spike Residue 403 Affects Binding of Coronavirus Spikes to Human ACE2.” Nature Communications 12 (1): 6855. https://doi.org/10.1038/s41467-021-27180-0. https://www.nature.com/articles/s41467-021-27180-0.