The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was first reported in Wuhan, China in 2019 and is the causal agent of the ongoing coronavirus disease 2019 (COVID-19) pandemic. SARS-CoV-2 is highly infectious, virulent, and has claimed more than 5 million lives worldwide.
Scientists have stated that it is imperative to determine the effects of various mutations present in the VOCs on the host’s immune response elicited through either vaccination or natural infection. The Alpha, Gamma, and Delta variants contain a deletion at the open reading frame 8 region (ORF8), making them inactive.
Interestingly, between 2002-2003, the ORF8 was identified as a mutation hotspot in SARS-CoV. This mutation is associated with host adaptation and viral replication.
The SARS-CoV-2 Δ382 variant
In Singapore and Taiwan, SARS-CoV-2 variants with the 382-nucleotide (nt) deletion at the ORF8 regions has been identified and termed as Δ382 SARS-CoV-2. SARS-CoV-2 variants with a mutation at the ORF8 region have also been identified in Australia (138-nt deletion), Bangladesh (345-nt deletion), and Spain (52-nt deletion).
Although deletion mutation at ORF8 of SARS-CoV-2 is linked with a mild infection, it accounts for around 5% of infections worldwide. Therefore, understanding the mechanism behind the mild infection would provide better insights for improved management of SARS-CoV-2 transmission and therapeutics. Additionally, this finding would elucidate the reason behind the high transmissibility of dominant VOCs.
Although several in vitro studies have indicated that SARS-CoV-2 ORF8 downregulates major histocompatibility complex (MHC) I molecules and blocks the type I interferon (IFN) signaling pathway, the functional impacts of the ORF8 deletion on the cellular host immune response against SARS-CoV-2 is not clear.
To address this research gap and understand the molecular mechanisms of this natural genetic deletion, a comprehensive characterization of the whole blood transcriptomic profiles and adaptive immune responses between SARS-CoV-2 infected individuals with wildtype (WT) and Δ382 SARS-CoV-2 was recently carried out by scientists in Singapore. The research is published in the Journal of Clinical Immunology.
Study findings
In the current study, researchers performed a high-density ribonucleic acid (RNA)-sequencing (RNA-seq) showing distinctly different transcriptomic profiles between WT and Δ382 SARS-CoV-2 strains.
Transcriptomic profiles of patients infected with Δ382 induced more active cellular stress response and an upregulated eIF2 signaling. This finding is in line with previous studies that reported coronaviruses elicited cellular stress responses following infection by targeting unfolded protein response (UPR) pathways. As a result, these responses lead to an imbalance in cellular homeostasis and trigger endoplasmic reticulum (ER) stress by specifically targeting activating transcription factor 6 (ATF6) which, in turn, promotes viral replication.
The authors indicated that the SARS-CoV-2 ORF8 protein triggers the protein kinase RNA-like ER kinase (PERK) and eIF2 signaling mechanisms.
The researchers also reported an under-expression of neutrophil activation-associated signature in Δ382 SARS-CoV-2 infected patients. Additionally, strong evidence of effector cytotoxic genes, as well as upregulation of SARS-CoV-2 specific T-cell immunity and antibody responses were observed, all of which strongly indicated robust T- and B-cell responses.
Previous studies have also reported that the SARS-CoV-2 ORF8 interacts with MHC-I molecules and, subsequently, downregulates the cytotoxic functions of T lymphocytes. In the current study, the researchers found the presence of several cytotoxic effector genes such as GZMA, GZMB, ID2, and PLAC8 in Δ382 SARS-CoV-2 infected patients.
The researchers also analyzed the plasma of these patients and reported elevated levels of IFN-γ, tumor necrosis factor α (TNF-α), and interleukin 2 (IL-2) during the acute phase of virus infection. These findings agree with previous reports showing high effector populations with a cytotoxic phenotype that is associated with effector CD8+, mucosal-associated invariant T-cells (MAIT), and natural killer (NK) T-cells in COVID-19 patients with milder symptoms.
The current study also revealed that the deletion of ORF8 could result in increased immunogenicity against SARS-CoV-2. Interestingly, Δ382-infected individuals showed higher immunoglobulin G (IgG) levels at the early acute phase of infection. However, more studies are required for a complete understanding of the roles of IgG in SARS-CoV-2 infection.
The researchers also found lower pro-inflammatory cytokines, chemokines, and growth factors in Δ382 SARS-CoV-2 infected patients. These factors are strongly associated with severe COVID-19. Pro-inflammatory Th1 responses were found to be more prominent in WT-infected patients.
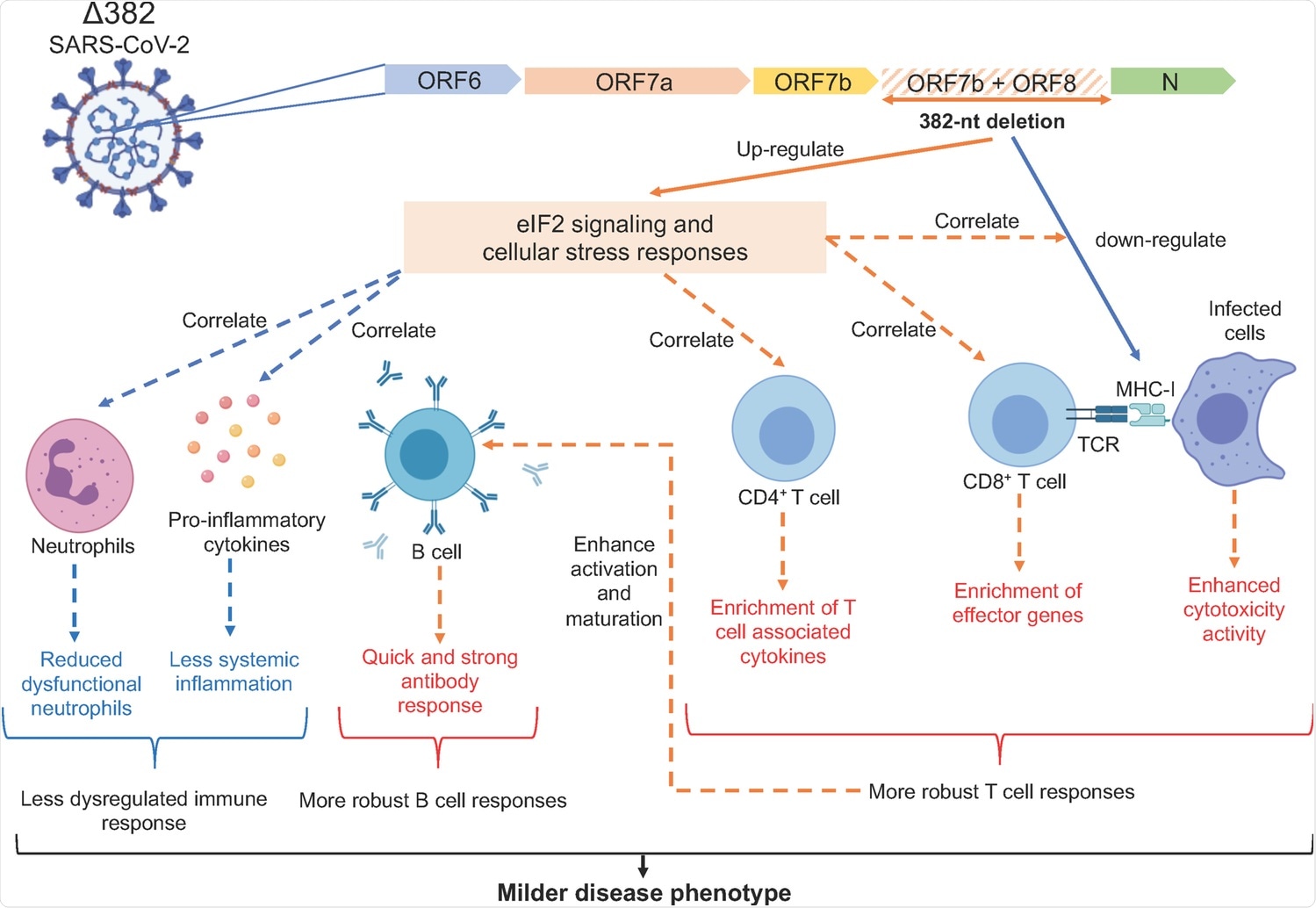 Molecular mechanisms underlying the milder disease phenotype in Δ382 SARS-CoV-2 infections. A SARS-CoV-2 variant with a 382-nucleotide deletion (Δ382) truncates ORF7b and removes the ORF8 transcription-regulatory sequence, eliminating ORF8 transcription. The ORF8 382-nt deletion has recently been associated with a milder disease phenotype. The attenuation of SARS-CoV-2 ORF8 upregulates eIF2 signaling and cellular stress responses at the acute phase of infection, potentially interrupting the downregulation of MHC-I molecules by ORF8 and also enhances the activation of both CD4+ and CD8+ T cells, evidenced by enrichment of effector cytotoxic genes and upregulation of SARS-CoV-2 specific T cell responses in Δ382 SARS-CoV-2 infected patients. Enhanced T cell responses may in turn mediate rapid and effective antibody responses in Δ382 SARS-CoV-2 infection. More pronounced cellular stress responses may further reduce systemic inflammation and dysfunctional neutrophils in Δ382 SARS-CoV-2 infected patients. Overall, the attenuation of SARS-CoV-2 ORF8 produced a molecular phenotype characterized by more pronounced cellular stress responses and a less dysregulated immune phenotype with more robust T and B cell responses.
Molecular mechanisms underlying the milder disease phenotype in Δ382 SARS-CoV-2 infections. A SARS-CoV-2 variant with a 382-nucleotide deletion (Δ382) truncates ORF7b and removes the ORF8 transcription-regulatory sequence, eliminating ORF8 transcription. The ORF8 382-nt deletion has recently been associated with a milder disease phenotype. The attenuation of SARS-CoV-2 ORF8 upregulates eIF2 signaling and cellular stress responses at the acute phase of infection, potentially interrupting the downregulation of MHC-I molecules by ORF8 and also enhances the activation of both CD4+ and CD8+ T cells, evidenced by enrichment of effector cytotoxic genes and upregulation of SARS-CoV-2 specific T cell responses in Δ382 SARS-CoV-2 infected patients. Enhanced T cell responses may in turn mediate rapid and effective antibody responses in Δ382 SARS-CoV-2 infection. More pronounced cellular stress responses may further reduce systemic inflammation and dysfunctional neutrophils in Δ382 SARS-CoV-2 infected patients. Overall, the attenuation of SARS-CoV-2 ORF8 produced a molecular phenotype characterized by more pronounced cellular stress responses and a less dysregulated immune phenotype with more robust T and B cell responses.
Conclusion
Some of the limitations of this study include a limited number of Δ382 SARS-CoV-2 infections for analysis. Additionally, the host's genetic background and demographic data were not evaluated.
However, the main finding of this study is the functional implication of ORF8 on host immune surveillance. This indicates the utilization of ORF8 as a possible target for COVID-19 therapeutics. However, the rapid evolution of the ORF8 gene due to mutation compromises its suitability as an antiviral target.
In the future, research associated with engineered viruses and animal models could further elucidate the interactions and mechanisms between ORF8 and ER stress or T-cell responses during SARS-CoV-2 infection.