SARS-CoV-2 ORF7a protein deletions (del) have been reported since the initial coronavirus disease 2019 (COVID-19) pandemic period. Delta outbreaks occurred in Australia from June 2021 to February 2022, and whole-genome sequencing (WGS) of the causative Delta variant showed ORF7a truncation due to the presence of a 17-nucleotide (nt) frameshift-inducing deletion.
 Study: Persistence of a SARS-CoV-2 variant with a frameshifting deletion for the duration of a major outbreak. Image Credit: Dotted Yeti / Shutterstock
Study: Persistence of a SARS-CoV-2 variant with a frameshifting deletion for the duration of a major outbreak. Image Credit: Dotted Yeti / Shutterstock

 *Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
About the study
In the present retrospective study, researchers investigated the persistence of ORF7aΔ17del for months, unlike other mutations in ORF7a that have been detected worldwide in Delta genomes. They investigated if Delta-ORF7aΔ17del produced a detectable/functional protein, the site of ORF7a localization in infected cells and assessed ORF7aΔ17del growth in vitro.
All GK glade genomes (predominantly comprising Delta genomes) were downloaded from the GISAID (global initiative on sharing all influenza) database. Each genome in the dataset was searched for ORF7a del by aligning each sequence against the Wuhan Hu-1 reference SARS-CoV-2 genome. Subsequently, genomes were discarded if they were not assigned to Delta, were duplicates, did not cover the entire ORF7a gene, and/or the collection dates were not mentioned.
Structural modeling and protein docking analyses were performed for ORF7a. Further, Calu-3 (lung carcinoma cell line) cells were infected with 103 TCID50 (50% tissue culture infectious dose) of A.2.2, Delta-ORF7aintact, or Delta-ORF7aΔ17del and western blot analysis was performed for cell supernatants of the infected and uninfected cells. In addition, immunofluorescence analysis was performed to assess the in vitro viral growth, and the mean fluorescent intensity (MFI) staining and expression patterns of ORF7a in A.2.2-infected and Delta-ORF7aΔ17del-infected cells were compared.
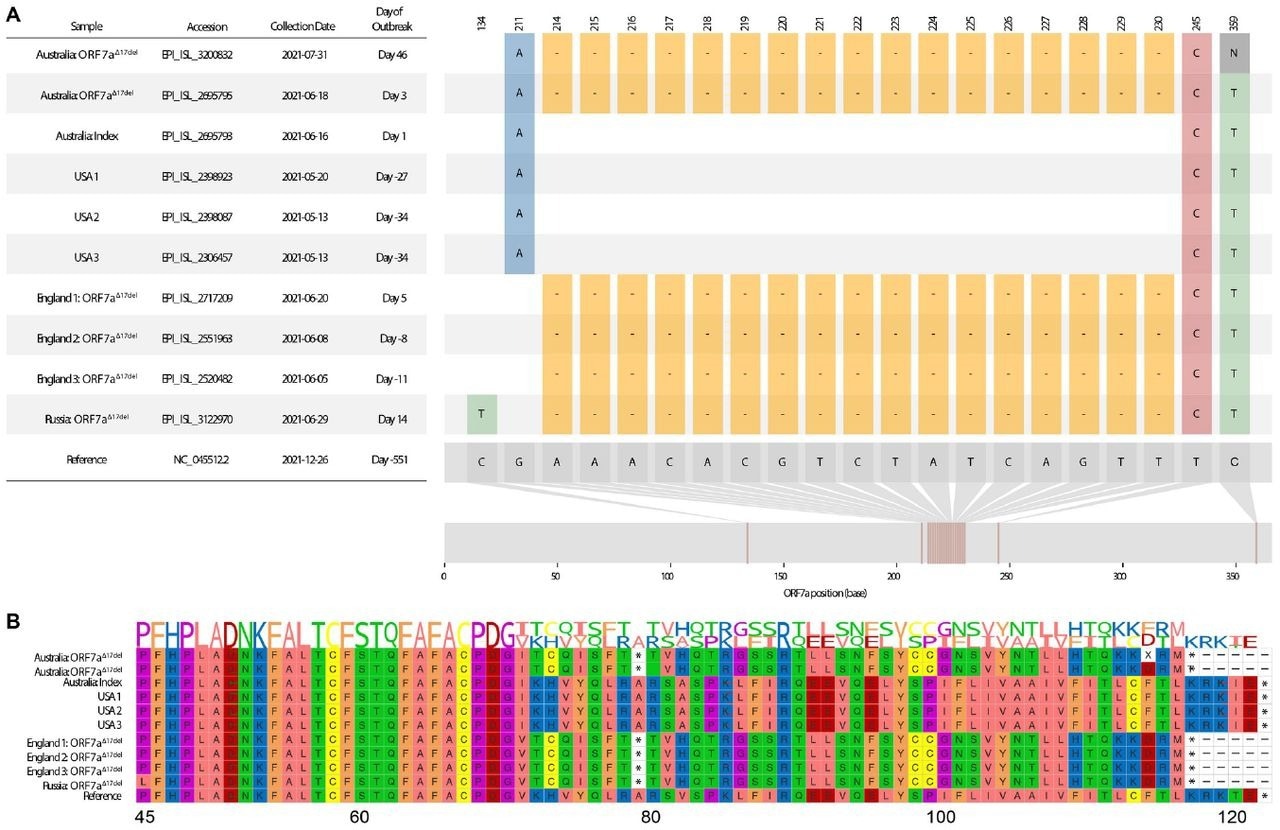 Visualization of a 17-nucleotide deletion in ORF7a of Delta variant SARS-CoV-2 sequences (ORF7aΔ17del), and the corresponding consequences at the peptide level. Sequences chosen for visualization include the index case of the Australian Delta outbreak (lacking ORF7aΔ17del), the first Australian case possessing ORF7aΔ17del during the outbreak period, an Australian sample from ∼1.5 months into the outbreak that possesses ORF7a 17del, the three USA samples (all lacking ORF7aΔ17del) that match the Australian index case, and a selection of four non-Australian samples to possess ORF7aΔ17del, some of which were collected before the Australian outbreak had begun. A. Sample dates of all chosen samples relative to the index case of the Australian Delta outbreak (16 June 2021), and all variants in the sample genomes relative to the SARS-CoV-2 reference genome (created in part using snipit (https://github.com/aineniamh/snipit). B. The consequences of ORF7aΔ17del on the amino acid translation of ORF7a, leading to a premature stop codon after amino acid 78 and a truncated peptide sequence (∼64% complete). Amino acids 1–44 are identical to the SARS-CoV-2 reference genome in all samples and are not presented.
Visualization of a 17-nucleotide deletion in ORF7a of Delta variant SARS-CoV-2 sequences (ORF7aΔ17del), and the corresponding consequences at the peptide level. Sequences chosen for visualization include the index case of the Australian Delta outbreak (lacking ORF7aΔ17del), the first Australian case possessing ORF7aΔ17del during the outbreak period, an Australian sample from ∼1.5 months into the outbreak that possesses ORF7a 17del, the three USA samples (all lacking ORF7aΔ17del) that match the Australian index case, and a selection of four non-Australian samples to possess ORF7aΔ17del, some of which were collected before the Australian outbreak had begun. A. Sample dates of all chosen samples relative to the index case of the Australian Delta outbreak (16 June 2021), and all variants in the sample genomes relative to the SARS-CoV-2 reference genome (created in part using snipit (https://github.com/aineniamh/snipit). B. The consequences of ORF7aΔ17del on the amino acid translation of ORF7a, leading to a premature stop codon after amino acid 78 and a truncated peptide sequence (∼64% complete). Amino acids 1–44 are identical to the SARS-CoV-2 reference genome in all samples and are not presented.
Results
A total of 4,018,216 Delta genomes were downloaded from the GISAID database, of which 4,195 distinct patterns were identified, and 3.4% (n=134,751) contained an ORF7a deletion, the most common being ORF7aΔ17del mutation. Almost all (99%) of Delta-ORF7aΔ17del genomes originated from the Australian outbreak and comprised 87% (27912 out of 32048) of the genomes of the outbreak.
In vitro experiments showed a considerably higher proportion of Delta-ORF7aΔ17del-infected cells compared to those infected with Delta-ORF7aintact, and A.2.2, indicative of a marginal growth advantage for Delta-ORF7aΔ17del over Delta-ORF7aintact. Delta-ORF7aΔ17del strains produced ORF7a, although in significantly lesser amounts compared to that by A.2.2, with a more diffuse pattern of expression throughout the cytoplasm of infected cells.
Over 70% of ORF7a del were estimated to cause a frameshift which would lead to a preterm stop codon and a truncated sequence of amino acids (aa). The del were usually identified over short periods (median of one day), with the most persistent in-frame del being a 12-nt del of 27617-27628 bases among 99 genomes, sampled between 11 August and last 12 January 2022, with a 519-day time difference. Frameshift-inducing del were usually transient; however, a few could be sampled for an extended period, e.g., a 64-nt deletion spanning over 27556-27619 positions was detected in 8945 sequences that were sampled between 20 March 2021 and 22 February 2022, with a 339-day time difference.
The most commonly detected ORF7a del corresponded to positions between 27607 and 27623, as identified in the Delta-ORF7aΔ17del genomes. The 17-nt del of Delta-ORF7aΔ17del occurred after the sixth β(beta)-sheet, causing loss of the C-terminal transmembrane domain, seventh β sheet, and the ER localization determining-cytoplasmic KRKTE motif.
Additionally, it led to ORF7a:K119 aa loss and inhibition of interferon type I (IFN-I) via signal transducer and activator of transcription 2 (STAT2) blockage. The ORF7a:K119 aa loss was estimated to lower Delta-ORF7aΔ17del variant replication titers. The 17 nt-del and N-terminal peptide retention could increase ORF7a secretion and alter Delta-ORF7aΔ17del variant pathology since ORF7a is usually retained on the cellular surfaces and intracellularly in the ER membrane and Golgi apparatus. Delta-ORF7aΔ17del variant’s ORF7A protein could bind with the cluster of differentiation 14 (CD14); however, the seventh β sheet and K119 loss caused significant changes in the interaction regions.
SARS-CoV-2 spike (S) and nucleocapsid (N) proteins were detected in all variants’ cellular supernatants but not from uninfected cells. About 79%, 61%, and 67% of cells infected with Delta-ORF7aΔ17del, Delta-ORF7aintact, and A.2.2, were SARS-CoV-2 N-positive, respectively, whereas no uninfected cells showed SARS-CoV-2 N positivity.
MFI staining of ORF7a among cells infected with Delta-ORF7aΔ17del and A.2.2 were 70.7 and 74.4, respectively, and ORF7a was undetectable among uninfected cells. The main probabilities explaining Delta-ORF7aΔ17del persistence in Australia were (i) no competition from Delta-ORF7aintact and other circulating variants, and (ii) the ORF7aΔ17del mutation may not be deleterious to Delta.
Overall, the study findings showed that Delta-ORF7aΔ17del genome proliferation during the Australian Delta outbreak most likely occurred due to chance founder effects rather than an intrinsic benefit of the ORF7aΔ17del mutation. Nevertheless, ORF7a deletion abundance in genomes globally indicates some potential benefits to Delta transmission. In addition, the findings indicated ORF7a as a hotspot for SARS-CoV-2 genome del; however, further genomic epidemiological research is warranted.

 *Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
Journal reference:
- Preliminary scientific report.
Persistence of a SARS-CoV-2 variant with a frameshifting deletion for the duration of a major outbreak. Charles S.P. Foster, Rowena A. Bull, Nicodemus Tedla, Fernando Santiago, David Agapiou, Anurag Adhikari, Gregory J. Walker, Lok Bahadur Shrestha, Sebastiaan J. van Hal, Ki Wook Kim, William D. Rawlinson, medRxiv preprint 2022, DOI: https://doi.org/10.1101/2022.09.06.22279658, https://www.medrxiv.org/content/10.1101/2022.09.06.22279658v1