The coronavirus disease 2019 (COVID-19) pandemic is caused by the highly infectious virus named severe acute respiratory syndrome coronavirus 2 (SARS‑CoV‑2). The SARS-CoV-2 pandemic still wreaks havoc on medical facilities worldwide despite the availability of vaccinations and antivirals against COVID-19. Therefore, robust techniques to screen for the presence of the SARS-CoV-2 in animal models or infected cells are a must for studying the basic biology of COVID-19 and the effects of antiviral therapies.
Recombinant viruses that express reporters and are replication-competent have previously been proved as a great way to research topics like SARS-CoV-2 infection, pathogenesis, replication, and transmission. Despite the establishment and efficient synthesis of rSARS-CoV-2 carrying fluorescent or luciferase reporter genes, the knowledge gained from their application across in vitro experiments or live animals was restricted to the properties of the fluorescent or luciferase reporter genes.
About the study
In the present research, the authors, for the first time, developed a replication-competent rSARS-CoV-2 that expresses both luciferase (Nluc) and fluorescent (mCherry) reporter genes (rSARS-CoV-2/mCherry-Nluc) to circumvent the drawbacks of using a single reporter gene.
The team developed the first rSARS-CoV-2 that expresses both the luciferase Nluc and fluorescence mCherry reporter genes, i.e., rSARS-CoV-2/mCherry-Nluc, using their previously disclosed bacterial artificial chromosome (BAC)-based reverse genetics and the innovative 2A method. The expression of the two reporter genes was confirmed using luciferase activity with a microplate reader (Nluc) and fluorescence microscopy (mCherry). The researchers used Western blotting to validate reporter expression.
The investigators created a bireporter-based microneutralization experiment to recognize and classify antivirals and neutralizing antibodies (NAbs). This assay was based on the benefits of using an rSARS-CoV-2 expressing both fluorescent and bioluminescence proteins versus those expressing either fluorescence or luciferase reporter genes. They infected keratin 18 human angiotensin-converting enzyme 2 (K18-hACE2) transgenic mice and golden Syrian hamsters with rSARS-CoV-2/mCherry-Nluc to see if the expression of mCherry coupled with Nluc altered SARS-CoV-2 replication in vivo.
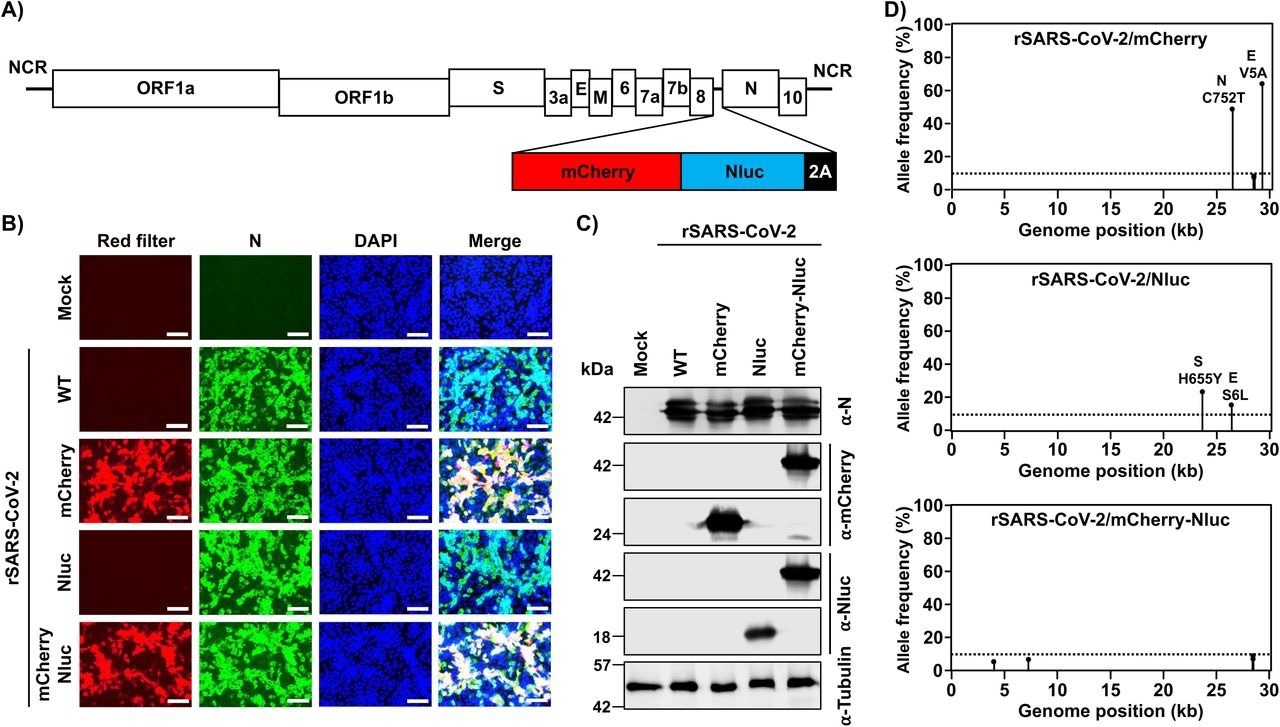
Generation of a bireporter rSARS-CoV-2 expressing mCherry and Nluc (rSARS-CoV-2/mCherry-Nluc). A) Schematic representation of the rSARS-CoV-2/mCherry-Nluc viral genome: SARS-CoV-2 structural, non-structural, and accessory open reading frame (ORF) proteins are indicated in white boxes. mCherry (red), Nluc (blue) and the PTV-1 2A autoproteolytic sequence (black) were inserted in front of the viral N protein. B) mCherry expression and immunofluorescence assays: Vero E6 cells (1.2 x 106 cells/well, 6-well format, triplicates) were mock-infected or infected (MOI 0.01) with rSARS-CoV-2 WT, rSARS-CoV-2/mCherry, rSARS-CoV-2/Nluc, or rSARS-CoV-2/mCherry-Nluc. Cells were fixed in 10% neutral buffered formalin 24 hpi before directly visualizing mCherry expression under a fluorescence microscope or the viral N protein using a specific 1C7C7 MAb. Cell nuclei were strained with DAPI. Representative images are shown. Scale bars = 100 µm. Magnification = X20. C) Western blots: Vero E6 cells (1.2 x 106 cells/well, 6-well format, triplicates) were mock-infected or infected (MOI 0.01) with rSARS-CoV-2 WT, rSARS-CoV-2/mCherry, rSARS-CoV-2/Nluc, or rSARS-CoV-2/mCherry-Nluc. At 24 hpi, cells were collected and protein expression in cell lysates were evaluated by Western blot using specific antibodies against SARS-CoV-2 N protein, or the mCherry and Nluc reporter proteins. Tubulin was included as a loading control. The molecular mass of proteins is indicated in kilodaltons (kDa) on the left. D) Deep sequencing analysis of reporter-expressing rSARS-CoV-2: The non-reference allele frequency of rSARS-CoV-2/mCherry (top), rSARS-CoV-2/Nluc (middle), and rSARS-CoV-2/mCherry-Nluc (bottom) was calculated by comparing the short reads to the respective reference SARS-CoV-2 WA-1 viral genome (MN985325.1). Non-reference alleles present in less than 10% of reads are not shown (dotted line) and the non-reference allele frequency that is greater than 10% is indicated.
Results
The study results illustrated that rSARS-CoV-2/mCherry-Nluc showcased identical viral fitness in cultured cells to rSARS-CoV-2 expressing single reporter luciferase (rSARS-CoV-2/Nluc) and fluorescent (rSARS-CoV-2/mCherry) genes or wild-type (WT) rSARS-CoV-2, which was an rSARS-CoV-2 without reporter genes. rSARS-CoV-2/mCherry-Nluc also harbored similar expression degrees for the two reporter genes.
Similarly, the new rSARS-CoV-2/mCherry-Nluc plaque phenotype was comparable in size to the plaque phenotypes of rSARS-CoV-2/Nluc, rSARS-CoV-2/mCherry, or rSARS-CoV-2/WT. However, just rSARS-CoV-2/mCherry-Nluc possessed measurable levels of expression of both reporter genes. The hypothesis that reporter genes were a reliable surrogate to investigate viral infection was further supported by the finding that Nluc or mCherry reporter expression levels coincided with degrees of viral replication.
The authors discovered that the half-maximal effective concentration (EC50) of antivirals and the 50% neutralizing titer (NT50) of NAbs achieved in bireporter-based microneutralization experiments using either fluorescence or luciferase signal were identical to those obtained with rSARS-CoV-2/WT, rSARS-CoV-2/Nluc or rSARS-CoV-2/mCherry reporter genes, and those documented in the prior literature.
In vivo, rSARS-CoV-2/mCherry-Nluc, in comparison to rSARS-CoV-2 harboring individual reporter genes or WT rSARS-CoV-2, exhibits similar pathogenicity in K18-hACE2 transgenic mice. Identical findings were reported in the golden Syrian hamster model of SARS-CoV-2 transmission and infection. Furthermore, in golden Syrian hamsters, rSARS-CoV-2/mCherry-Nluc permits the evaluation of COVID-19 and SARS-CoV-2 transmission using in vivo imaging systems (IVIS).
Taken together, the current study shows that it was possible to analyze SARS-CoV-2 in vivo and in vitro utilizing the presently established novel bireporter-expressing rSARS-CoV-2, i.e., rSARS-CoV-2/mCherry-Nluc.
Conclusions
On the whole, the team created an rSARS-CoV-2 that expresses the luciferase (Nluc) and fluorescent (mCherry) genes, namely rSARS-CoV-2/mCherry-Nluc, in the present study. They showed that rSARS-CoV-2/mCherry-Nluc could be used to research SARS-CoV-2 biology in vivo and or in vitro, such as characterization and identification of NAbs and or antivirals. The scientists depict SARS-CoV-2 infection and transmission via IVIS using rodent models.
rSARS-CoV-2/mCherry-Nluc was appropriate for various experimental applications presently unavailable by using rSARS-CoV-2 expressing a single luciferase or fluorescent reporter gene. According to the investigators, this SARS-CoV-2/mCherry-Nluc virus was the initial replication-competent rSARS-CoV-2 consistently expressing two reporter genes.
The viability of producing rSARS-CoV-2 expressing a combination of two reporter genes illustrates the flexibility of the viral genome to express extensive open reading frames (ORFs) from the locus of the SARS-CoV-2 nucleocapsid (N) protein. Besides, the ability to express foreign genes without having to remove a viral protein (like ORF7a) and the robust thresholds of reporter gene expression acquired employing 2A autoproteolytic approach make rSARS704 CoV-2/mCherry-Nluc an excellent choice for studying viral infection, transmission, and pathogenesis, such as newly discovered SARS-CoV-2 variants of concern (VOCs).

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Monitoring SARS-CoV-2 infection using a double reporter-expressing virus; Kevin Chiem, Jun-Gyu Park, Desarey Morales Vasquez, Richard K. Plemper, Jordi B Torrelles, James Kobie, Mark R Walter, Chengjin Ye, Luis Martinez-Sobrido. bioRxiv preprint 2022, DOI: https://doi.org/10.1101/2022.06.23.497376, https://www.biorxiv.org/content/10.1101/2022.06.23.497376v1
- Peer reviewed and published scientific report.
Chiem, Kevin, Jun-Gyu Park, Desarey Morales Vasquez, Richard K. Plemper, Jordi B. Torrelles, James J. Kobie, Mark R. Walter, Chengjin Ye, and Luis Martinez-Sobrido. 2022. “Monitoring SARS-CoV-2 Infection Using a Double Reporter-Expressing Virus.” Edited by Daniela S. Rajao. Microbiology Spectrum 10 (5). https://doi.org/10.1128/spectrum.02379-22. https://journals.asm.org/doi/10.1128/spectrum.02379-22.