In response to the changing SARS-CoV-2, researchers have created a new vaccine targeting its S2 subunit. This strategy focuses on stabilizing the subunit's prefusion state, enhancing the immune response. Maintaining the subunit structure and antigenicity of S2, that vaccine proves effective against different sarbecovirus clades. It elicited robust antibody responses in animal tests, chiefly against difficult-to-resolve variants such as XBB.1.5. While promising, further research is required to confirm its efficacy in humans and its long-term effectiveness against new SARS-CoV-2 variants.
About the study
The present study employed various cell lines, including Human Embryonic Kidney 293 cells with SV40 T-antigen (HEK293T), Expi293F, and VeroE6-TMPRSS2. These cells were cultivated in specific media conditions, though not authenticated or tested for mycoplasma contamination. The focus was on producing recombinant S2 antigen proteins. Utilizing Expi293F cells, deoxyribonucleic acid (DNA) transfections were carried out, followed by a precise harvesting process. The proteins were then purified using specialized techniques and equipment, ensuring their quality for further experiments.
HexaPro S glycoproteins of both the SARS-CoV-2 and the SARS-CoV-1 were produced following specific protocols in Expi293F cells. After transfection, the proteins underwent a similar purification technique to maintain their stability and usability.
Monoclonal antibody Enzyme-linked immunosorbent assays (ELISAs) were conducted using carefully prepared proteins and specific protocols to determine EC50 values. Meanwhile, HEK293T cells were used to produce vesicular stomatitis virus (VSV) pseudoviruses expressing various S constructs. This time-consuming process went through several steps to maintain the quality and effectiveness of pseudoviruses.
Serological ELISAs were also carried out, employing a systematic approach to analyze the immunogenicity of various proteins. Additionally, the study utilized negative stain electron microscopy and cryo-electron microscopy for detailed structural analysis, following rigorous preparation and data collection protocols. The immunogenicity aspect was thoroughly examined through experiments on BALB/c mice, following specific guidelines and protocols. This included detailed immunization schedules and serum collection for comprehensive analysis.
Finally, neutralization assays were performed using VeroE6-TMPRSS2 cells and various pseudoviruses. This involved a series of well-orchestrated steps to accurately measure the neutralization capacity of the sera, providing critical insights into the effectiveness of the immunogens. The data from these assays were carefully analyzed to determine ID50 values, contributing to the overall understanding of the vaccine candidates' potential.
Study results
The research team previously designed a fusion machinery (S2 subunit) antigen stabilized by introducing specific HexaPro mutations, an inter-protomer disulfide, and an intra-protomer disulfide. This construct, named C-44, exhibited a prefusion tertiary structure but a splayed-open quaternary structure. To achieve a native quaternary structure in the prefusion S2 subunit trimer, they selected mutations from a deep-mutational scanning dataset, focusing on prefusion-stabilizing amino acid substitutions. Individual mutations, namely T961F, D994E, and Q1005R, were introduced into the C-44 background and recombinantly produced. These constructs showed monodispersity and high yields of purified protein.
Electron microscopy (EM) characterization revealed that the T961F mutation (E-31) formed closed S2 trimers, unlike other constructs which adopted various conformations. CryoEM structure determination at 2.7 Å resolution confirmed the prefusion closed structure of E-31, with the T961F substitution reinforcing interactions at the fusion machinery apex. Although a fraction of E-31 trimers with an open apex was detected, the mutation effectively closed the apex in a prefusion conformation.
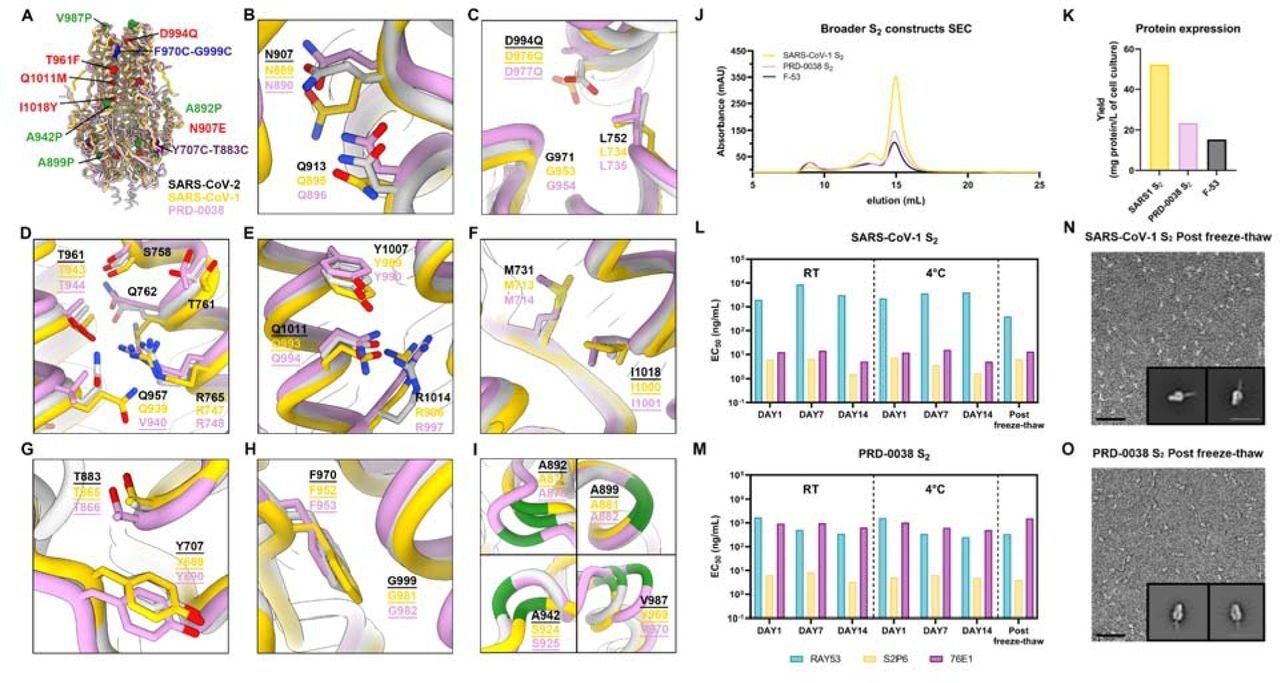
A broadly generalizable prefusion-stabilization strategy for sarbecovirus fusion machinery (S2 subunit) antigens. A, Ribbon diagrams of superimposed S2 subunits of the prefusion SARS-CoV-2 S (PDB 6VXX), SARS-CoV-1 S (PDB 5X58) and PRD-0038 S (PDB 8U29) structures. Prefusion-stabilizing mutations are shown in blue (intra-protomer disulfide bond), purple (VFLIP inter-protomer disulfide bond), red (mutations ported from E-69), and green (subset of proline mutations selected from HexaPro). B-I, Zoomed-in views of superimposed S2 subunits of the prefusion SARS-CoV-2, SARS-CoV-1 and PRD-0038 S structures highlighting the local structural conservation of residues mutated in SARS-CoV-2 E-69/F-53. Mutated residues in our designed constructs are underlined. SARS-CoV-2, SARS-CoV-1, and PRD-0038 S are respectively shown in light gray, gold, and pink in panels (A-I). J, Size-exclusion chromatograms of the designed SARS-CoV-1 and PRD-0038 S2 constructs, as compared to SARS-CoV-2 F53. K, Purification yields of the designed SARS-CoV-1 and PRD-0038 S2 constructs. The yield for the best SARS-CoV-2 S2 construct (F53) is included for comparison. L,M, Evaluation of retention of antigenicity for the SARS-CoV-1 (L) and PRD-0038 (M) S2 antigens in various storage conditions using binding of the S2P6, 76E1 and RAY53 monoclonal antibodies analyzed by ELISA. N,O, Evaluation of retention of the native prefusion conformation of the negatively stained SARS-CoV-1 (N) and PRD-0038 (O) S2 trimers after freeze/thawing. Insets: 2D class averages showing compact prefusion S2 trimers. The scale bar represents 50 nm (micrographs) and 200 Å (2D class averages).
To improve conformational homogeneity, additional constructs with more mutations were designed. The E-60 construct contained nine additional mutations, but its cryoEM structure indicated distortion in an α-helix region. A new construct, E-69, was designed with four additional mutations, resulting in a prefusion closed S2 subunit trimer without open apex trimers, as confirmed by a 3Å resolution cryoEM structure.
Antigenicity retention was tested under various storage conditions to assess the stability of the E-69 design. ELISA and EM analyses showed that E-69 retained its prefusion conformation and unaltered antigenicity, even after freezing and thawing. This stability suggested validity in the prefusion-stabilization strategy, making E-69 a promising vaccine candidate.
The broad applicability of the S2 subunit prefusion-stabilization strategy was then evaluated across different sarbecovirus clades. Constructs were designed for SARS-CoV-1 S2 and PRD-0038 S2, incorporating mutations from E-69. These constructs showed high yields and stability, retaining their antigenicity and adopting the intended closed prefusion architecture.
The immunogenicity of the E-69 vaccine candidate was tested in BALB/c mice with various vaccination schedules. ELISA analyses showed comparable antibody binding titers against SARS-CoV-2 variants and higher titers against SARS-CoV-1. Neutralizing antibody titers were highest in E-69-vaccinated mice against the XBB.1.5 variant, demonstrating the potential of this vaccine to elicit broadly reactive antibody responses.
Finally, to assess the in vivo protective efficacy, mice were challenged with the XBB.1.5 variant. While none of the vaccines prevented infection, vaccinated mice were protected against weight loss and had comparable viral titers, suggesting the potential of the S2 subunit vaccine to protect against immune evasive SARS-CoV-2 variants.

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.